













| General Information: |
|
| Name(s) found: |
KAR2 /
YJL034W
[SGD]
|
| Description(s) found:
Found 34 descriptions. SHOW ALL |
|
| Organism: | Saccharomyces cerevisiae |
| Length: | 682 amino acids |
Gene Ontology: |
|
| Cellular Component: |
endoplasmic reticulum
[IDA |
| Biological Process: |
SRP-dependent cotranslational protein targeting to membrane, translocation
[IMP karyogamy involved in conjugation with cellular fusion [IGI ER-associated protein catabolic process [IMP posttranslational protein targeting to membrane, translocation [IDA response to unfolded protein [IMP |
| Molecular Function: |
ATPase activity
[IDA unfolded protein binding [IMP |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MFFNRLSAGK LLVPLSVVLY ALFVVILPLQ NSFHSSNVLV RGADDVENYG TVIGIDLGTT 60
61 YSCVAVMKNG KTEILANEQG NRITPSYVAF TDDERLIGDA AKNQVAANPQ NTIFDIKRLI 120
121 GLKYNDRSVQ KDIKHLPFNV VNKDGKPAVE VSVKGEKKVF TPEEISGMIL GKMKQIAEDY 180
181 LGTKVTHAVV TVPAYFNDAQ RQATKDAGTI AGLNVLRIVN EPTAAAIAYG LDKSDKEHQI 240
241 IVYDLGGGTF DVSLLSIENG VFEVQATSGD THLGGEDFDY KIVRQLIKAF KKKHGIDVSD 300
301 NNKALAKLKR EAEKAKRALS SQMSTRIEID SFVDGIDLSE TLTRAKFEEL NLDLFKKTLK 360
361 PVEKVLQDSG LEKKDVDDIV LVGGSTRIPK VQQLLESYFD GKKASKGINP DEAVAYGAAV 420
421 QAGVLSGEEG VEDIVLLDVN ALTLGIETTG GVMTPLIKRN TAIPTKKSQI FSTAVDNQPT 480
481 VMIKVYEGER AMSKDNNLLG KFELTGIPPA PRGVPQIEVT FALDANGILK VSATDKGTGK 540
541 SESITITNDK GRLTQEEIDR MVEEAEKFAS EDASIKAKVE SRNKLENYAH SLKNQVNGDL 600
601 GEKLEEEDKE TLLDAANDVL EWLDDNFETA IAEDFDEKFE SLSKVAYPIT SKLYGGADGS 660
661 GAADYDDEDE DDDGDYFEHD EL |
| PUBLICATION | TOPOLOGY | COCOMPLEXED PROTEINS | |
| View Details | Hazbun TR, et al. (2003) |
|
GRS1 KAR2 TUB3 YNL313C |
| View Details | Krogan NJ, et al. (2006) |
|
EMP24 ERP1 ERP2 KAR2 PHB2 SCJ1 SSS1 YEL1 |
| View Details | Gavin AC, et al. (2006) |
|
KAR2 SCJ1 VPS52 VPS53 VPS54 |
The following runs contain data for this protein:
SHOWING SINGLE HITS. [ Hide Single Hits ]
| BAIT | PREY | HITS | PUBLICATION | |
| View Screen | 1 | Unpublished Fields Lab Data | ||
| View Screen | 1 | Unpublished Fields Lab Data | ||
| View Screen | 1 | Unpublished Fields Lab Data |
New Feature: Upload Your Own Microscopy Data
| PROTEIN(S) | PUBLICATION | |
| View Data |
|
Huh WK, et al. (2003) |
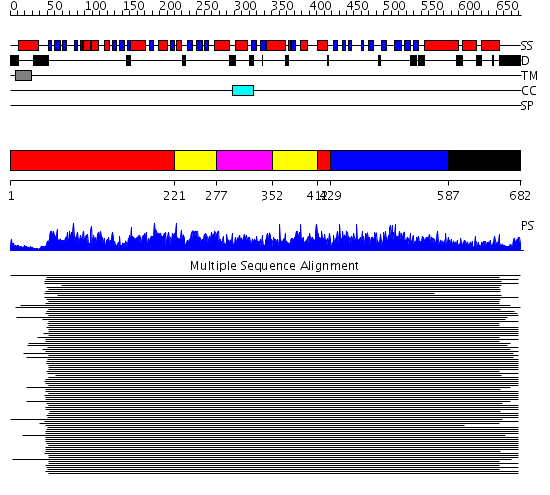
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..220] [412..428] |
2450.0 | Heat shock protein 70kDa, ATPase fragment | |
| 2 | View Details | [221..276] [352..411] |
2450.0 | Heat shock protein 70kDa, ATPase fragment | |
| 3 | View Details | [277..351] | 2450.0 | Heat shock protein 70kDa, ATPase fragment | |
| 4 | View Details | [429..586] | 668.201249 | DnaK | |
| 5 | View Details | [587..682] | 3.0 | Heavy meromyosin subfragment |
Functions predicted (by domain):
| # | Gene Ontology predictions | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 1 |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 2 | No functions predicted. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 3 | No functions predicted. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 4 | No functions predicted. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 5 | No functions predicted. |
| Source: Reynolds et al. 2008. Manuscript submitted |

|