













| General Information: |
|
| Name(s) found: |
PRE4 /
YFR050C
[SGD]
|
| Description(s) found:
Found 36 descriptions. SHOW ALL |
|
| Organism: | Saccharomyces cerevisiae |
| Length: | 266 amino acids |
Gene Ontology: |
|
| Cellular Component: |
proteasome core complex, beta-subunit complex
[IPI |
| Biological Process: |
ubiquitin-dependent protein catabolic process
[TAS |
| Molecular Function: |
endopeptidase activity
[TAS |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MNHDPFSWGR PADSTYGAYN TQIANAGASP MVNTQQPIVT GTSVISMKYD NGVIIAADNL 60
61 GSYGSLLRFN GVERLIPVGD NTVVGISGDI SDMQHIERLL KDLVTENAYD NPLADAEEAL 120
121 EPSYIFEYLA TVMYQRRSKM NPLWNAIIVA GVQSNGDQFL RYVNLLGVTY SSPTLATGFG 180
181 AHMANPLLRK VVDRESDIPK TTVQVAEEAI VNAMRVLYYR DARSSRNFSL AIIDKNTGLT 240
241 FKKNLQVENM KWDFAKDIKG YGTQKI |
The following runs contain data for this protein:
| BAIT | COMMENTS | PUBLICATION | |
| View Run | sample cbf3 from feb 2005 | Sandall S, et all (2006) | |
| View Run | Sample 2- cpn2 from june 2005 | McCusker D, et al (2007) | |
| View Run | Sample boi 1 with ha tag from october 2005 | McCusker D, et al (2007) | |
| View Run | #12 Mitotic Prep1-TiO2 Flowthrough | Keck JM, et al. (2011) |
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
New Feature: Upload Your Own Microscopy Data
| PROTEIN(S) | PUBLICATION | |
| View Data |
|
Huh WK, et al. (2003) |
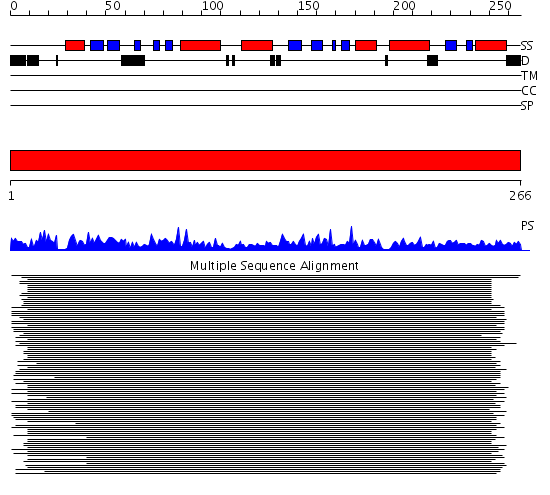
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..266] | 1563.0 | Proteasome beta subunit (catalytic) |
Functions predicted (by domain):
| # | Gene Ontology predictions | ||||||||||||||||||||||||||||||
| 1 |
|
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.92 |
Source: Reynolds et al. (2008)