













| General Information: |
|
| Name(s) found: |
HRT1 /
YOL133W
[SGD]
|
| Description(s) found:
Found 29 descriptions. SHOW ALL |
|
| Organism: | Saccharomyces cerevisiae |
| Length: | 121 amino acids |
Gene Ontology: |
|
| Cellular Component: |
nucleus
[IDA nuclear ubiquitin ligase complex [TAS cytoplasm [IDA SCF ubiquitin ligase complex [IDA Cul3-RING ubiquitin ligase complex [NAS |
| Biological Process: |
G2/M transition of mitotic cell cycle
[TAS G1/S transition of mitotic cell cycle [IGI SCF-dependent proteasomal ubiquitin-dependent protein catabolic process [IDA protein ubiquitination during ubiquitin-dependent protein catabolic process [IDA |
| Molecular Function: |
ubiquitin-protein ligase activity
[IDA |
| PUBLICATION | TOPOLOGY | COCOMPLEXED PROTEINS | |
| View Details | Riffle et al. (2010) (Unpublished Data) |

|
CDC34 CDC53 HRT1 |
| View Details | Krogan NJ, et al. (2006) |
|
CLG1 CUE1 GTR1 HAM1 HRT1 LAP3 TFP1 VMA10 VMA13 VMA2 VMA7 VMA8 XDJ1 YIG1 |
| View Details | Gavin AC, et al. (2002) |

|
BRR1 CLF1 CSN12 CUS1 CWC2 DIB1 ECM2 HRT1 HSH155 HSH49 LEA1 LSM4 LUC7 MSL1 MUD1 PRP11 PRP19 PRP21 PRP3 PRP31 PRP39 PRP4 PRP40 PRP42 PRP43 PRP46 PRP6 PRP9 RSE1 SMB1 SMD1 SMD2 SMD3 SME1 SMX2 SNP1 SNT309 SNU114 SNU56 SNU66 SNU71 SPP382 SRB2 STO1 |
| View Details | Ho Y, et al. (2002) |

|
ADR1 BBC1 CDC39 CDC53 CRM1 DUR1,2 ECM29 ECM33 FAA4 GAL3 GCN1 GUF1 HRT1 HYP2 IDH1 MDN1 MET30 MKT1 MMS1 MYO2 PFK1 PMA2 RIX7 RPA190 RPN1 RPN8 RTT101 SEC27 SKP1 TPS1 UBI4 VPS13 YAR009C YGP1 YLR035C-A |
| View Details | Qiu et al. (2008) |
|
APC1 CDC34 CDC4 CDC53 GRR1 HRT1 MET30 SGT1 SKP1 UFO1 |
SHOWING SINGLE HITS. [ Hide Single Hits ]
New Feature: Upload Your Own Microscopy Data
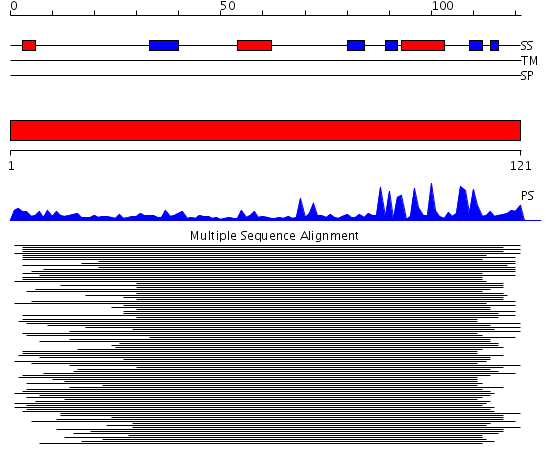
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..121] | 309.30103 | RIGG-box protein 1 (RBX1) of SCF ubiquitin ligase complex |
Functions predicted (by domain):
| # | Gene Ontology predictions | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 1 |
|
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.99 |
Source: Reynolds et al. (2008)