













| General Information: |
|
| Name(s) found: |
BBC1 /
YJL020C
[SGD]
|
| Description(s) found:
Found 23 descriptions. SHOW ALL |
|
| Organism: | Saccharomyces cerevisiae |
| Length: | 1157 amino acids |
Gene Ontology: |
|
| Cellular Component: |
actin cortical patch
[IDA |
| Biological Process: |
actin cytoskeleton organization
[IGI |
| Molecular Function: |
myosin I binding
[IDA |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MSEPEVPFKV VAQFPYKSDY EDDLNFEKDQ EIIVTSVEDA EWYFGEYQDS NGDVIEGIFP 60
61 KSFVAVQGSE VGKEAESSPN TGSTEQRTIQ PEVEQKDLPE PISPETKKET LSGPVPVPAA 120
121 TVPVPAATVP VPAATAVSAQ VQHDSSSGNG ERKVPMDSPK LKARLSMFNQ DITEQVPLPK 180
181 STHLDLENIP VKKTIVADAP KYYVPPGIPT NDTSNLERKK SLKENEKKIV PEPINRAQVE 240
241 SGRIETENDQ LKKDLPQMSL KERIALLQEQ QRLQAAREEE LLRKKAKLEQ EHERSAVNKN 300
301 EPYTETEEAE ENEKTEPKPE FTPETEHNEE PQMELLAHKE ITKTSREADE GTNDIEKEQF 360
361 LDEYTKENQK VEESQADEAR GENVAEESEI GYGHEDREGD NDEEKEEEDS EENRRAALRE 420
421 RMAKLSGASR FGAPVGFNPF GMASGVGNKP SEEPKKKQHK EKEEEEPEQL QELPRAIPVM 480
481 PFVDPSSNPF FRKSNLSEKN QPTETKTLDP HATTEHEQKQ EHGTHAYHNL AAVDNAHPEY 540
541 SDHDSDEDTD DHEFEDANDG LRKHSMVEQA FQIGNNESEN VNSGEKIYPQ EPPISHRTAE 600
601 VSHDIENSSQ NTTGNVLPVS SPQTRVARNG SINSLTKSIS GENRRKSINE YHDTVSTNSS 660
661 ALTETAQDIS MAAPAAPVLS KVSHPEDKVP PHPVPSAPSA PPVPSAPSVP SAPPVPPAPP 720
721 ALSAPSVPPV PPVPPVSSAP PALSAPSIPP VPPTPPAPPA PPAPLALPKH NEVEEHVKSS 780
781 APLPPVSEEY HPMPNTAPPL PRAPPVPPAT FEFDSEPTAT HSHTAPSPPP HQNVTASTPS 840
841 MMSTQQRVPT SVLSGAEKES RTLPPHVPSL TNRPVDSFHE SDTTPKVASI RRSTTHDVGE 900
901 ISNNVKIEFN AQERWWINKS APPAISNLKL NFLMEIDDHF ISKRLHQKWV VRDFYFLFEN 960
961 YSQLRFSLTF NSTSPEKTVT TLQERFPSPV ETQSARILDE YAQRFNAKVV EKSHSLINSH1020
1021 IGAKNFVSQI VSEFKDEVIQ PIGARTFGAT ILSYKPEEGI EQLMKSLQKI KPGDILVIRK1080
1081 AKFEAHKKIG KNEIINVGMD SAAPYSSVVT DYDFTKNKFR VIENHEGKII QNSYKLSHMK1140
1141 SGKLKVFRIV ARGYVGW |
| PUBLICATION | TOPOLOGY | COCOMPLEXED PROTEINS | |
| View Details | Krogan NJ, et al. (2006) |
|
BBC1 PPR1 SKT5 UBP7 |
| View Details | Gavin AC, et al. (2006) |

|
BBC1 MTC1 |
| View Details | Ho Y, et al. (2002) |
|
ADR1 BBC1 CDC39 CDC53 CPR1 CRM1 DUR1,2 ECM29 ECM33 FAA4 GAL3 GCN1 GSY2 HRT1 HTB2 HYP2 MDN1 MKT1 MMS1 MYO2 PFK1 PMA2 RPA190 RPN1 TPS1 UBI4 VPS13 YAR009C YGP1 YLR035C-A |
| View Details | Qiu et al. (2008) |

|
ABP1 AIP1 ARP2 BBC1 BNI1 CAP1 CAP2 COF1 CRN1 END3 LAS17 PAN1 PFY1 RVS167 SAC6 SCP1 SLA1 SRV2 TPM2 TWF1 VRP1 |
The following runs contain data for this protein:
| BAIT | COMMENTS | PUBLICATION | |
| View Run | Dam1-765 complex purified from yeast with Dad1-TAP | Shimogawa MM, et al. (2006) | |
| View Run | sample cbf3 from feb 2005 | Sandall S, et all (2006) | |
| View Run | No Comments | Schneider, DA, et al. (2006) | |
| View Run | Sample 2- cpn2 from june 2005 | McCusker D, et al (2007) | |
| View Run | Sample 3 - cpn + atp, from june 2005 | McCusker D, et al (2007) | |
| View Run | Sample boi 1 with ha tag from october 2005 | McCusker D, et al (2007) | |
| View Run | #22 Asynchronous Prep2-IMAC Phosphopeptide enrichment B1-2 | Keck JM, et al. (2011) | |
| View Run | #29 Asynchronous Prep5-TiO2 Flowthrough | Keck JM, et al. (2011) |
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
New Feature: Upload Your Own Microscopy Data
| PROTEIN(S) | PUBLICATION | |
| View Data |
|
Huh WK, et al. (2003) |
| View Data |
|
Huh WK, et al. (2003) |
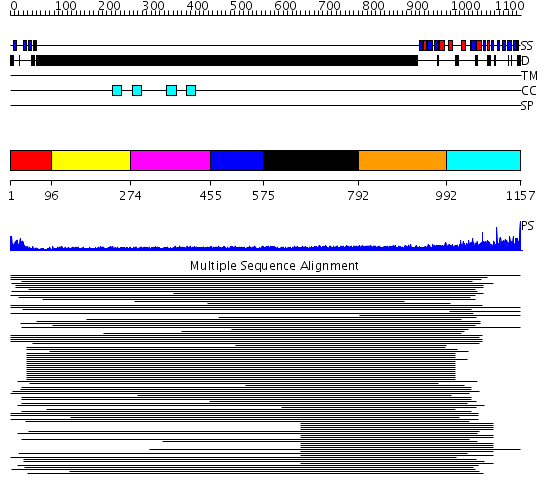
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..95] | 6.69897 | Hemapoetic cell kinase Hck; Hemopoetic cell kinase Hck; Haemopoetic cell kinase Hck | |
| 2 | View Details | [96..273] | N/A | No confident structure predictions are available. | |
| 3 | View Details | [274..454] | N/A | No confident structure predictions are available. | |
| 4 | View Details | [455..574] | N/A | No confident structure predictions are available. | |
| 5 | View Details | [575..791] | N/A | No confident structure predictions are available. | |
| 6 | View Details | [792..991] | 2.305994 | View MSA. No confident structure predictions are available. | |
| 7 | View Details | [992..1157] | 2.264996 | View MSA. No confident structure predictions are available. |
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.99 |
Source: Reynolds et al. (2008)