













| General Information: |
|
| Name(s) found: |
PSA1 /
YDL055C
[SGD]
|
| Description(s) found:
Found 32 descriptions. SHOW ALL |
|
| Organism: | Saccharomyces cerevisiae |
| Length: | 361 amino acids |
Gene Ontology: |
|
| Cellular Component: |
cytoplasm
[IDA |
| Biological Process: |
cell wall mannoprotein biosynthetic process
[IMP GDP-mannose biosynthetic process [IDA protein amino acid glycosylation [IMP |
| Molecular Function: |
mannose-1-phosphate guanylyltransferase activity
[IDA |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MKGLILVGGY GTRLRPLTLT VPKPLVEFGN RPMILHQIEA LANAGVTDIV LAVNYRPEVM 60
61 VETLKKYEKE YGVNITFSVE TEPLGTAGPL KLAEDVLKKD NSPFFVLNSD VICEYPFKEL 120
121 ADFHKAHGGK GTIVATKVDE PSKYGVIVHD IATPNLIDRF VEKPKEFVGN RINAGLYILN 180
181 PEVIDLIEMK PTSIEKETFP ILVEEKQLYS FDLEGFWMDV GQPKDFLSGT VLYLNSLAKR 240
241 QPKKLATGAN IVGNALIDPT AKISSTAKIG PDVVIGPNVT IGDGVRITRS VVLCNSTIKN 300
301 HSLVKSTIVG WNSTVGQWCR LEGVTVLGDD VEVKDEIYIN GGKVLPHKSI SDNVPKEAII 360
361 M |
| PUBLICATION | TOPOLOGY | COCOMPLEXED PROTEINS | |
| View Details | Riffle et al. (2010) (Unpublished Data) |
|
GDH2 PGA2 PSA1 SYP1 YNR062C |
| View Details | Krogan NJ, et al. (2006) |
|
CDC31 HOS3 PSA1 PTK2 RPL1A, RPL1B |
| View Details | Gavin AC, et al. (2006) |
|
ILV1 PSA1 YOL098C |
| View Details | Gavin AC, et al. (2002) |

|
ACC1 CCT8 CLU1 CPA2 FDH1 GCN20 GFA1 GUS1 ILV1 KAP123 LYS12 MRPL3 PSA1 RPA135 RPB2 RPN10 RPN11 RPN12 RPN8 RPT3 RPT5 RPT6 SAM1 SEC27 SMC3 TCB1 TIF1, TIF2 TSA1 TUB3 URA7 VAS1 YOL098C |
The following runs contain data for this protein:
| BAIT | COMMENTS | PUBLICATION | |
| View Run | No Comments | Reinke A, et al. (2004) | |
| View Run | No Comments | Reinke A, et al. (2004) | |
| View Run | No Comments | De Craene, J., et al. (2006) | |
| View Run | No Comments | Widlund PO, et al. (2005) | |
| View Run | Dam1 complex purified from yeast with Dad1-TAP | Shimogawa MM, et al. (2006) | |
| View Run | his-HA tag on RPA135 | Schneider, DA, et al. (2006) | |
| View Run | his-HA tag on RPA135 | Schneider, DA, et al. (2006) | |
| View Run | Dam1-765 complex purified from yeast with Dad1-TAP | Shimogawa MM, et al. (2006) | |
| View Run | sample bir 1 from feb 2005 | Sandall S, et all (2006) | |
| View Run | Dam1-765 complex purified from yeast with Dad1-TAP | Shimogawa MM, et al. (2006) | |
| View Run | Dam1 complex purified from yeast with Dad1-TAP | Shimogawa MM, et al. (2006) | |
| View Run | Sample 2- cpn2 from june 2005 | McCusker D, et al (2007) | |
| View Run | Sample 3 - cpn + atp, from june 2005 | McCusker D, et al (2007) | |
| View Run | Sample: HX11 - both fusion and trans-SNARE complex formation take place. | Hao Xu, et al. (2010) | |
| View Run | Sample: HX12 - trans-SNARE complex formation and fusion are inhibited (control). | Hao Xu, et al. (2010) | |
| View Run | Sample: HX14 - mixture in detergent (control). | Hao Xu, et al. (2010) | |
| View Run | #25 Asynchronous Prep3-TiO2 Flowthrough | Keck JM, et al. (2011) | |
| View Run | #24 Asynchronous Prep3-TiO2 Phosphopeptide enrichment | Keck JM, et al. (2011) | |
| View Run | #34 Asynchronous SPB prep | Keck JM, et al. (2011) | |
| View Run | #32 Asynchronous Prep-No Phosphopeptide enrichment | Keck JM, et al. (2011) | |
| View Run | #31 Asynchronous Prep (Protease cleavage) | Keck JM, et al. (2011) | |
| View Run | #08 Alpha Factor Prep4-TiO2 Phosphopeptide enrichment | Keck JM, et al. (2011) |
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
New Feature: Upload Your Own Microscopy Data
| PROTEIN(S) | PUBLICATION | |
| View Data |
|
Huh WK, et al. (2003) |
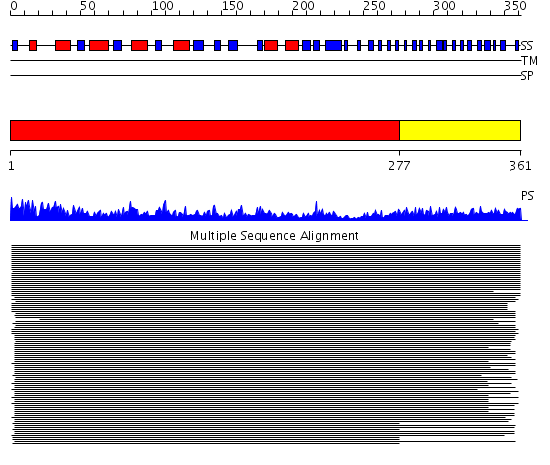
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..276] | 238.66807 | RmlA (RfbA) | |
| 2 | View Details | [277..361] | 3.0 | gamma-carbonic anhydrase |
Functions predicted (by domain):
| # | Gene Ontology predictions | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 1 |
|
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 2 | No functions predicted. |
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.98 |
Source: Reynolds et al. (2008)