| |
Size |
Notes |
Members |
| View GO Analysis |
22 proteins |
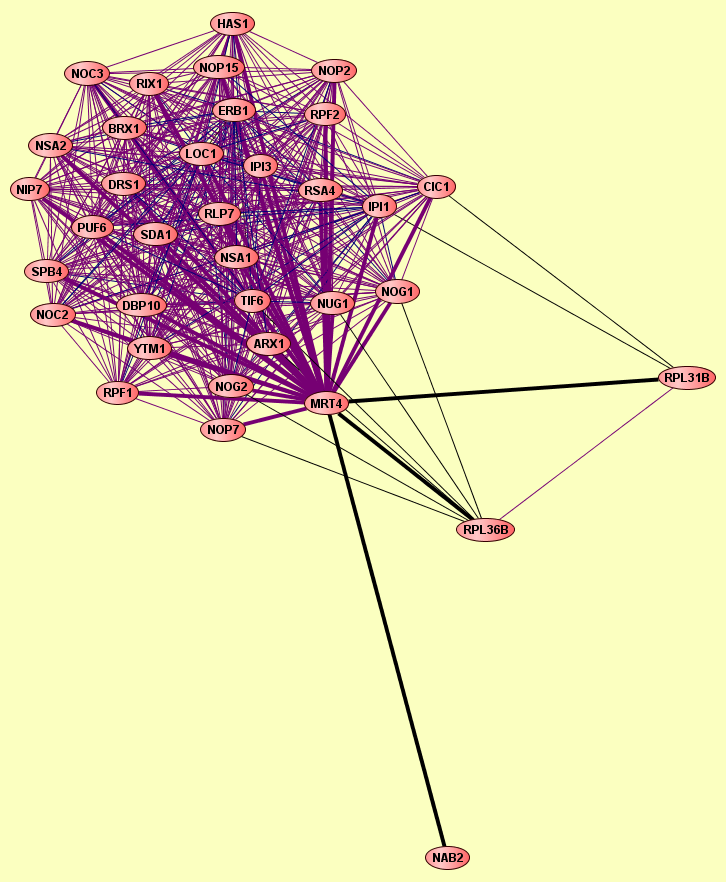
Complex predicted from the combined set of Gavin (2002), Gavin (2006), Ho (2002) and Krogan (2006); at p-value cutoff of 1E-7 |
BRX1
CIC1
DBP10
DRS1
ERB1
HAS1
MRT4
NIP7
NOC2
NOC3
NOG1
NOP15
NOP2
NOP7
NSA1
NUG1
PUF6
RIX1
RLP7
RPF1
SPB4
YTM1
|
| View GO Analysis |
15 proteins |
Complex predicted from the combined set of Gavin (2002), Gavin (2006), Ho (2002) and Krogan (2006); at p-value cutoff of 1E-7 |
BRX1
CIC1
DBP10
DRS1
HAS1
MRT4
NOC2
NOC3
NOG1
NOP15
NOP2
NOP7
NUG1
PUF6
RPF2
|
| View GO Analysis |
22 proteins |
Complex predicted from the combined set of Gavin (2002), Gavin (2006), Ho (2002) and Krogan (2006); at p-value cutoff of 1E-7 |
CIC1
DBP10
DRS1
ERB1
HAS1
MRT4
NIP7
NOC2
NOC3
NOG1
NOP15
NOP2
NOP7
NSA1
NUG1
PUF6
RIX1
RLP7
RPF1
SPB4
TIF6
YTM1
|
| View GO Analysis |
13 proteins |
Complex predicted from the combined set of Gavin (2002), Gavin (2006), Ho (2002) and Krogan (2006); at p-value cutoff of 1E-7 |
ARX1
IPI3
MRT4
NOG1
NOG2
NOP7
NSA2
NUG1
RIX1
RLP7
RSA4
SDA1
TIF6
|
| View GO Analysis |
15 proteins |
Complex predicted from the combined set of Gavin (2002), Gavin (2006), Ho (2002) and Krogan (2006); at p-value cutoff of 1E-7 |
CIC1
DBP10
DRS1
HAS1
MRT4
NOC2
NOC3
NOG1
NOP15
NOP2
NOP7
NUG1
PUF6
RPF2
TIF6
|
| View GO Analysis |
13 proteins |
Complex predicted from the combined set of Gavin (2002), Gavin (2006), Ho (2002) and Krogan (2006); at p-value cutoff of 1E-7 |
CIC1
DBP10
HAS1
MRT4
NOG1
NOG2
NOP15
NOP7
NSA2
NUG1
PUF6
RPF2
TIF6
|
| View GO Analysis |
13 proteins |
Complex predicted from the combined set of Gavin (2002), Gavin (2006), Ho (2002) and Krogan (2006); at p-value cutoff of 1E-7 |
CIC1
DBP10
HAS1
MRT4
NOC3
NOG1
NOG2
NOP15
NOP7
NUG1
PUF6
RPF2
TIF6
|
| View GO Analysis |
14 proteins |
Complex predicted from the combined set of Gavin (2002), Gavin (2006), Ho (2002) and Krogan (2006); at p-value cutoff of 1E-7 |
CIC1
DBP10
DRS1
HAS1
MRT4
NOC2
NOG1
NOP15
NOP7
NSA2
NUG1
PUF6
RPF2
TIF6
|
| View GO Analysis |
19 proteins |
Complex predicted from the combined set of Gavin (2002), Gavin (2006), Ho (2002) and Krogan (2006); at p-value cutoff of 1E-7 |
CIC1
DBP10
DRS1
ERB1
HAS1
MRT4
NOC2
NOG1
NOP15
NOP7
NSA1
NSA2
NUG1
PUF6
RIX1
RLP7
RPF1
TIF6
YTM1
|
| View GO Analysis |
16 proteins |
Complex predicted from the combined set of Gavin (2002), Gavin (2006), Ho (2002) and Krogan (2006); at p-value cutoff of 1E-7 |
ARX1
CIC1
DBP10
ERB1
HAS1
MRT4
NIP7
NOC3
NOG1
NOG2
NOP15
NOP7
NUG1
RIX1
RLP7
TIF6
|
| View GO Analysis |
17 proteins |
Complex predicted from the combined set of Gavin (2002), Gavin (2006), Ho (2002) and Krogan (2006); at p-value cutoff of 1E-7 |
ARX1
CIC1
DBP10
DRS1
ERB1
HAS1
MRT4
NIP7
NOC3
NOG1
NOP15
NOP7
NSA1
NUG1
RIX1
RLP7
TIF6
|
| View GO Analysis |
16 proteins |
Complex predicted from the combined set of Gavin (2002), Gavin (2006), Ho (2002) and Krogan (2006); at p-value cutoff of 1E-7 |
CIC1
DBP10
ERB1
HAS1
MRT4
NIP7
NOC3
NOG1
NOG2
NOP15
NOP7
NUG1
PUF6
RIX1
RLP7
TIF6
|
| View GO Analysis |
17 proteins |
Complex predicted from the combined set of Gavin (2002), Gavin (2006), Ho (2002) and Krogan (2006); at p-value cutoff of 1E-7 |
CIC1
DBP10
DRS1
ERB1
HAS1
MRT4
NOG1
NOP15
NOP7
NSA2
NUG1
RIX1
RLP7
RPF1
SDA1
TIF6
YTM1
|
| View GO Analysis |
16 proteins |
Complex predicted from the combined set of Gavin (2002), Gavin (2006), Ho (2002) and Krogan (2006); at p-value cutoff of 1E-7 |
ARX1
CIC1
DBP10
ERB1
HAS1
MRT4
NOG1
NOG2
NOP15
NOP7
NSA2
NUG1
RIX1
RLP7
SDA1
TIF6
|
| View GO Analysis |
15 proteins |
Complex predicted from the combined set of Gavin (2002), Gavin (2006), Ho (2002) and Krogan (2006); at p-value cutoff of 1E-7 |
ARX1
CIC1
DBP10
IPI3
MRT4
NOG1
NOG2
NOP15
NOP7
NSA2
NUG1
RIX1
RLP7
SDA1
TIF6
|
| View GO Analysis |
16 proteins |
Complex predicted from the combined set of Gavin (2002), Gavin (2006), Ho (2002) and Krogan (2006); at p-value cutoff of 1E-7 |
ARX1
CIC1
DBP10
DRS1
ERB1
HAS1
MRT4
NOG1
NOP15
NOP7
NSA2
NUG1
RIX1
RLP7
SDA1
TIF6
|
| View GO Analysis |
15 proteins |
Complex predicted from the combined set of Gavin (2002), Gavin (2006), Ho (2002) and Krogan (2006); at p-value cutoff of 1E-7 |
CIC1
DBP10
ERB1
HAS1
MRT4
NOG1
NOG2
NOP15
NOP7
NSA2
NUG1
PUF6
RIX1
RLP7
TIF6
|
| View GO Analysis |
16 proteins |
Complex predicted from the combined set of Gavin (2002), Gavin (2006), Ho (2002) and Krogan (2006); at p-value cutoff of 1E-7 |
ARX1
CIC1
DBP10
DRS1
ERB1
HAS1
MRT4
NOG1
NOP15
NOP7
NSA1
NSA2
NUG1
RIX1
RLP7
TIF6
|
| View GO Analysis |
14 proteins |
Complex predicted from the combined set of Gavin (2002), Gavin (2006), Ho (2002) and Krogan (2006); at p-value cutoff of 1E-7 |
CIC1
DBP10
IPI3
MRT4
NOG1
NOP15
NOP7
NSA2
NUG1
RIX1
RLP7
RPF1
SDA1
TIF6
|
| View GO Analysis |
8 proteins |
Complex predicted from the combined set of Gavin (2002), Gavin (2006), Ho (2002) and Krogan (2006); at p-value cutoff of 1E-7 |
ARX1
MRT4
NOG1
NOG2
NOP7
NUG1
RPL36B
TIF6
|
| View GO Analysis |
10 proteins |
Complex predicted from the combined set of Gavin (2002), Gavin (2006), Ho (2002) and Krogan (2006); at p-value cutoff of 1E-7 |
ARX1
IPI1
IPI3
MRT4
NOG2
NOP7
NUG1
RIX1
RSA4
SDA1
|
| View GO Analysis |
2 proteins |
Complex predicted from the combined set of Gavin (2002), Gavin (2006), Ho (2002) and Krogan (2006); at p-value cutoff of 1E-7 |
MRT4
RPL31B
|
| View GO Analysis |
9 proteins |
Complex predicted from the combined set of Gavin (2002), Gavin (2006), Ho (2002) and Krogan (2006); at p-value cutoff of 1E-7 |
CIC1
DBP10
DRS1
HAS1
LOC1
MRT4
NOP2
NUG1
RPF2
|
| View GO Analysis |
10 proteins |
Complex predicted from the combined set of Gavin (2002), Gavin (2006), Ho (2002) and Krogan (2006); at p-value cutoff of 1E-7 |
CIC1
DBP10
DRS1
HAS1
LOC1
MRT4
NOP2
NUG1
RPF1
YTM1
|
| View GO Analysis |
2 proteins |
Complex predicted from the combined set of Gavin (2002), Gavin (2006), Ho (2002) and Krogan (2006); at p-value cutoff of 1E-7 |
MRT4
NAB2
|