













| General Information: |
|
| Name(s) found: |
EFT1 /
YOR133W
[SGD]
EFT2 / YDR385W [SGD] |
| Description(s) found:
Found 70 descriptions. SHOW ALL |
|
| Organism: | Saccharomyces cerevisiae |
| Length: | 842 amino acids |
Gene Ontology: |
|
| Cellular Component: |
ribosome
[TAS |
| Biological Process: |
translational elongation
[TAS |
| Molecular Function: |
translation elongation factor activity
[TAS |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MVAFTVDQMR SLMDKVTNVR NMSVIAHVDH GKSTLTDSLV QRAGIISAAK AGEARFTDTR 60
61 KDEQERGITI KSTAISLYSE MSDEDVKEIK QKTDGNSFLI NLIDSPGHVD FSSEVTAALR 120
121 VTDGALVVVD TIEGVCVQTE TVLRQALGER IKPVVVINKV DRALLELQVS KEDLYQTFAR 180
181 TVESVNVIVS TYADEVLGDV QVYPARGTVA FGSGLHGWAF TIRQFATRYA KKFGVDKAKM 240
241 MDRLWGDSFF NPKTKKWTNK DTDAEGKPLE RAFNMFILDP IFRLFTAIMN FKKDEIPVLL 300
301 EKLEIVLKGD EKDLEGKALL KVVMRKFLPA ADALLEMIVL HLPSPVTAQA YRAEQLYEGP 360
361 ADDANCIAIK NCDPKADLML YVSKMVPTSD KGRFYAFGRV FAGTVKSGQK VRIQGPNYVP 420
421 GKKDDLFIKA IQRVVLMMGR FVEPIDDCPA GNIIGLVGID QFLLKTGTLT TSETAHNMKV 480
481 MKFSVSPVVQ VAVEVKNAND LPKLVEGLKR LSKSDPCVLT YMSESGEHIV AGTGELHLEI 540
541 CLQDLEHDHA GVPLKISPPV VAYRETVESE SSQTALSKSP NKHNRIYLKA EPIDEEVSLA 600
601 IENGIINPRD DFKARARIMA DDYGWDVTDA RKIWCFGPDG NGPNLVIDQT KAVQYLHEIK 660
661 DSVVAAFQWA TKEGPIFGEE MRSVRVNILD VTLHADAIHR GGGQIIPTMR RATYAGFLLA 720
721 DPKIQEPVFL VEIQCPEQAV GGIYSVLNKK RGQVVSEEQR PGTPLFTVKA YLPVNESFGF 780
781 TGELRQATGG QAFPQMVFDH WSTLGSDPLD PTSKAGEIVL AARKRHGMKE EVPGWQEYYD 840
841 KL |
| PUBLICATION | TOPOLOGY | COCOMPLEXED PROTEINS | |
| View Details | Riffle et al. (2010) (Unpublished Data) |
|
ADH1 ADH2 ADH3 CRN1 DPH1 DPH2 EFT2, EFT1 ENO1 FBA1 GCN20 GPM1 HSC82 HXK1 ILV1 KTI11 PDC1 RPA135 RPN2 SAC6 SAM1 SEC27 TIF1, TIF2 TKL1 YEF3 YHR020W |
| View Details | Krogan NJ, et al. (2006) |
|
AIM22 DPH1 DPH2 EFT2, EFT1 HGH1 MNN1 PGI1 RRG10 TRX1 YMR315W |
| View Details | Gavin AC, et al. (2006) |
|
DPH1 DPH2 EFT2, EFT1 HGH1 YOL098C |
| View Details | Qiu et al. (2008) |

|
EFT2, EFT1 RKI1 RPL10 RPL12B, RPL12A RPL13A RPL14A RPL14B RPL17B RPL1A, RPL1B RPL21A RPL21B RPL24A RPL26A RPL26B RPL27B RPL2A, RPL2B RPL32 RPL37A RPL37B RPL41A, RPL41B RPL42B, RPL42A RPL43B, RPL43A RPL5 RPL6A RPL9A RPL9B RPP1B RPP2A RPS12 RPS14B RPS16B, RPS16A RPS25A RPS27A RPS29A RPS29B RPS6A, RPS6B RPS7B RPS8B, RPS8A TEF1, TEF2 |
The following runs contain data for this protein:
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
New Feature: Upload Your Own Microscopy Data
| PROTEIN(S) | PUBLICATION | |
| View Data |
|
Huh WK, et al. (2003) |
| View Data |
|
Huh WK, et al. (2003) |
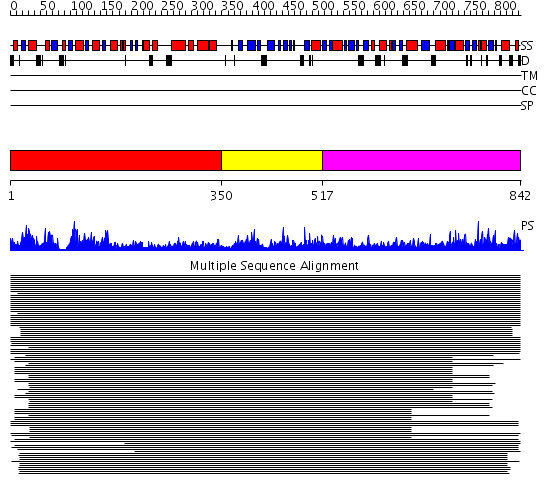
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..349] | 1300.9691 | Elongation factor G (EF-G), domain II; Elongation factor G (EF-G), N-terminal (G) domain; Elongation factor G (EF-G), domain IV; Elongation factor G (EF-G) | |
| 2 | View Details | [350..516] | 1300.9691 | Elongation factor G (EF-G), domain II; Elongation factor G (EF-G), N-terminal (G) domain; Elongation factor G (EF-G), domain IV; Elongation factor G (EF-G) | |
| 3 | View Details | [517..842] | 1300.9691 | Elongation factor G (EF-G), domain II; Elongation factor G (EF-G), N-terminal (G) domain; Elongation factor G (EF-G), domain IV; Elongation factor G (EF-G) |
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.99 |
Source: Reynolds et al. (2008)