| |
Size |
Notes |
Members |
| View GO Analysis |
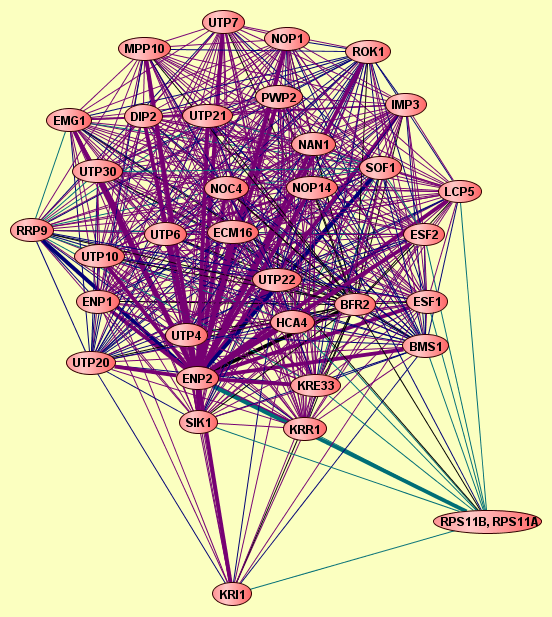
22 proteins |
Complex predicted from the combined set of Gavin (2002), Gavin (2006), Ho (2002) and Krogan (2006); at p-value cutoff of 1E-7 |
BMS1
DIP2
ECM16
EMG1
ENP2
IMP3
KRE33
MPP10
NAN1
NOC4
NOP1
NOP14
NOP56
PWP2
SOF1
UTP10
UTP20
UTP22
UTP30
UTP4
UTP6
UTP7
|
| View GO Analysis |
20 proteins |
Complex predicted from the combined set of Gavin (2002), Gavin (2006), Ho (2002) and Krogan (2006); at p-value cutoff of 1E-7 |
BMS1
ECM16
EMG1
ENP1
ENP2
IMP3
KRE33
MPP10
NAN1
NOC4
NOP1
NOP14
SOF1
UTP10
UTP20
UTP22
UTP30
UTP4
UTP6
UTP7
|
| View GO Analysis |
22 proteins |
Complex predicted from the combined set of Gavin (2002), Gavin (2006), Ho (2002) and Krogan (2006); at p-value cutoff of 1E-7 |
BMS1
DIP2
ECM16
EMG1
ENP2
IMP3
KRE33
MPP10
NAN1
NOC4
NOP1
NOP56
PWP2
RRP9
SOF1
UTP10
UTP20
UTP21
UTP30
UTP4
UTP6
UTP7
|
| View GO Analysis |
22 proteins |
Complex predicted from the combined set of Gavin (2002), Gavin (2006), Ho (2002) and Krogan (2006); at p-value cutoff of 1E-7 |
BMS1
DIP2
ECM16
EMG1
ENP2
IMP3
KRE33
MPP10
NAN1
NOC4
NOP1
NOP56
PWP2
SOF1
UTP10
UTP20
UTP21
UTP22
UTP30
UTP4
UTP6
UTP7
|
| View GO Analysis |
16 proteins |
Complex predicted from the combined set of Gavin (2002), Gavin (2006), Ho (2002) and Krogan (2006); at p-value cutoff of 1E-7 |
DIP2
EMG1
ENP2
IMP3
KRE33
MPP10
NAN1
NOC4
PWP2
ROK1
UTP10
UTP20
UTP21
UTP30
UTP6
UTP7
|
| View GO Analysis |
17 proteins |
Complex predicted from the combined set of Gavin (2002), Gavin (2006), Ho (2002) and Krogan (2006); at p-value cutoff of 1E-7 |
BFR2
BMS1
ECM16
EMG1
ENP1
ENP2
IMP3
KRE33
MPP10
NAN1
NOC4
NOP1
NOP14
UTP20
UTP30
UTP6
UTP7
|
| View GO Analysis |
16 proteins |
Complex predicted from the combined set of Gavin (2002), Gavin (2006), Ho (2002) and Krogan (2006); at p-value cutoff of 1E-7 |
BMS1
ECM16
EMG1
ENP1
ENP2
IMP3
KRE33
KRR1
MPP10
NOC4
NOP1
NOP14
SOF1
UTP20
UTP6
UTP7
|
| View GO Analysis |
16 proteins |
Complex predicted from the combined set of Gavin (2002), Gavin (2006), Ho (2002) and Krogan (2006); at p-value cutoff of 1E-7 |
BMS1
ECM16
EMG1
ENP2
IMP3
KRE33
KRR1
MPP10
NOC4
NOP1
NOP14
NOP56
SOF1
UTP20
UTP6
UTP7
|
| View GO Analysis |
17 proteins |
Complex predicted from the combined set of Gavin (2002), Gavin (2006), Ho (2002) and Krogan (2006); at p-value cutoff of 1E-7 |
BFR2
BMS1
ECM16
EMG1
ENP2
IMP3
KRE33
MPP10
NAN1
NOC4
NOP1
NOP14
NOP56
UTP20
UTP30
UTP6
UTP7
|
| View GO Analysis |
14 proteins |
Complex predicted from the combined set of Gavin (2002), Gavin (2006), Ho (2002) and Krogan (2006); at p-value cutoff of 1E-7 |
BFR2
EMG1
ENP2
IMP3
KRE33
MPP10
NAN1
NOC4
ROK1
UTP20
UTP21
UTP30
UTP6
UTP7
|
| View GO Analysis |
17 proteins |
Complex predicted from the combined set of Gavin (2002), Gavin (2006), Ho (2002) and Krogan (2006); at p-value cutoff of 1E-7 |
BFR2
BMS1
ECM16
EMG1
ENP2
IMP3
KRE33
MPP10
NAN1
NOC4
NOP1
NOP56
UTP20
UTP21
UTP30
UTP6
UTP7
|
| View GO Analysis |
7 proteins |
Complex predicted from the combined set of Gavin (2002), Gavin (2006), Ho (2002) and Krogan (2006); at p-value cutoff of 1E-7 |
BFR2
ENP2
ESF1
HCA4
KRI1
NOP56
RPS11A, RPS11B
|
| View GO Analysis |
6 proteins |
Complex predicted from the combined set of Gavin (2002), Gavin (2006), Ho (2002) and Krogan (2006); at p-value cutoff of 1E-7 |
BFR2
ENP2
ESF1
ESF2
HCA4
RPS11A, RPS11B
|
| View GO Analysis |
7 proteins |
Complex predicted from the combined set of Gavin (2002), Gavin (2006), Ho (2002) and Krogan (2006); at p-value cutoff of 1E-7 |
BFR2
ENP2
HCA4
KRE33
KRI1
NOP56
RPS11A, RPS11B
|
| View GO Analysis |
7 proteins |
Complex predicted from the combined set of Gavin (2002), Gavin (2006), Ho (2002) and Krogan (2006); at p-value cutoff of 1E-7 |
BFR2
BMS1
ENP2
ESF1
HCA4
NOP56
RPS11A, RPS11B
|
| View GO Analysis |
7 proteins |
Complex predicted from the combined set of Gavin (2002), Gavin (2006), Ho (2002) and Krogan (2006); at p-value cutoff of 1E-7 |
BFR2
BMS1
ENP2
HCA4
KRE33
NOP56
RPS11A, RPS11B
|
| View GO Analysis |
7 proteins |
Complex predicted from the combined set of Gavin (2002), Gavin (2006), Ho (2002) and Krogan (2006); at p-value cutoff of 1E-7 |
BMS1
ENP2
HCA4
KRE33
NOP56
RPS11A, RPS11B
UTP22
|
| View GO Analysis |
5 proteins |
Complex predicted from the combined set of Gavin (2002), Gavin (2006), Ho (2002) and Krogan (2006); at p-value cutoff of 1E-7 |
ENP2
HCA4
KRE33
RPS11A, RPS11B
UTP22
|
| View GO Analysis |
6 proteins |
Complex predicted from the combined set of Gavin (2002), Gavin (2006), Ho (2002) and Krogan (2006); at p-value cutoff of 1E-7 |
BFR2
ENP2
HCA4
KRE33
LCP5
RPS11A, RPS11B
|
| View GO Analysis |
6 proteins |
Complex predicted from the combined set of Gavin (2002), Gavin (2006), Ho (2002) and Krogan (2006); at p-value cutoff of 1E-7 |
ENP2
KRE33
KRI1
KRR1
NOP56
RPS11A, RPS11B
|
| View GO Analysis |
8 proteins |
Complex predicted from the combined set of Gavin (2002), Gavin (2006), Ho (2002) and Krogan (2006); at p-value cutoff of 1E-7 |
BMS1
ECM16
ENP2
KRE33
KRR1
NOP14
NOP56
RPS11A, RPS11B
|
| View GO Analysis |
8 proteins |
Complex predicted from the combined set of Gavin (2002), Gavin (2006), Ho (2002) and Krogan (2006); at p-value cutoff of 1E-7 |
BMS1
ECM16
ENP2
KRE33
NOP14
NOP56
RPS11A, RPS11B
UTP22
|
| View GO Analysis |
8 proteins |
Complex predicted from the combined set of Gavin (2002), Gavin (2006), Ho (2002) and Krogan (2006); at p-value cutoff of 1E-7 |
BFR2
BMS1
ECM16
ENP2
KRE33
NOP14
NOP56
RPS11A, RPS11B
|
| View GO Analysis |
5 proteins |
Complex predicted from the combined set of Gavin (2002), Gavin (2006), Ho (2002) and Krogan (2006); at p-value cutoff of 1E-7 |
ECM16
ENP2
KRE33
RPS11A, RPS11B
UTP22
|