













| General Information: |
|
| Name(s) found: |
QCR7 /
YDR529C
[SGD]
|
| Description(s) found:
Found 30 descriptions. SHOW ALL |
|
| Organism: | Saccharomyces cerevisiae |
| Length: | 127 amino acids |
Gene Ontology: |
|
| Cellular Component: |
mitochondrion
[IDA mitochondrial respiratory chain complex III [IDA |
| Biological Process: |
respiratory chain complex III assembly
[IMP mitochondrial electron transport, ubiquinol to cytochrome c [IMP aerobic respiration [IMP |
| Molecular Function: |
ubiquinol-cytochrome-c reductase activity
[IMP |
| PUBLICATION | TOPOLOGY | COCOMPLEXED PROTEINS | |
| View Details | (MIPS) Mewes HW, et al. (2004) |
|
COB COR1 CYT1 QCR10 QCR2 QCR6 QCR7 QCR8 QCR9 RIP1 |
| View Details | Ho Y, et al. (2002) |
|
QCR7 UBC4 UFD4 |
| View Details | Qiu et al. (2008) |

|
AAC1 ATP1 ATP14 ATP15 ATP16 ATP17 ATP18 ATP2 ATP20 ATP3 ATP4 ATP5 ATP6 ATP7 ATP8 COB COR1 COX1 COX12 COX13 COX2 COX3 COX4 COX5A COX5B COX6 COX7 COX8 COX9 CYC2 CYC7 CYT1 INH1 MCR1 OLI1 QCR10 QCR2 QCR6 QCR7 QCR8 QCR9 RIP1 SDH1 SDH2 SDH3 SDH4 |
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
New Feature: Upload Your Own Microscopy Data
| PROTEIN(S) | PUBLICATION | |
| View Data |
|
Huh WK, et al. (2003) |
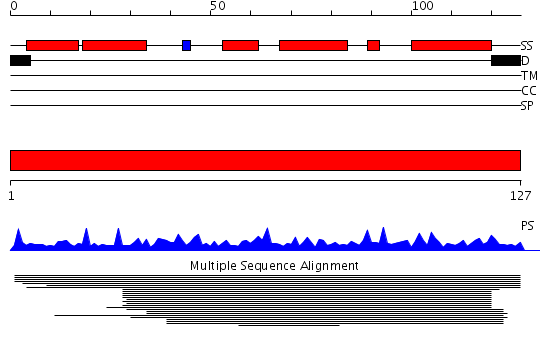
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..127] | 687.68867 | 14 kDa protein of cytochrome bc1 complex (Ubiquinol-cytochrome c reductase) |
Functions predicted (by domain):
| # | Gene Ontology predictions |
| 1 |
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.99 |
Source: Reynolds et al. (2008)