













| General Information: |
|
| Name(s) found: |
TAF2 /
YCR042C
[SGD]
|
| Description(s) found:
Found 26 descriptions. SHOW ALL |
|
| Organism: | Saccharomyces cerevisiae |
| Length: | 1407 amino acids |
Gene Ontology: |
|
| Cellular Component: |
transcription factor TFIID complex
[IDA |
| Biological Process: |
regulation of transcription involved in G1 phase of mitotic cell cycle
[IMP transcription initiation from RNA polymerase II promoter [TAS |
| Molecular Function: |
general RNA polymerase II transcription factor activity
[TAS |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MMSFSKNATP RAIVSESSTL HEMKFRNFRV AHEKISLDID LATHCITGSA TIIIIPLIQN 60
61 LEYVTFDCKE MTIKDVLVEN RRCDQFIHDD PLQTNLNGLT SQNVLYSDNS IEQSHFLRSK 120
121 FASLNEYPET DSKSQLTIKI PSSIKISLED ANALSNYTPI TPSIKTTPGF QESVFTPITL 180
181 QIEYEIRNPK SGIKFDTVYA DKPWLWNVYT SNGEICSSAS YWVPCVDLLD EKSTWELEFS 240
241 VPRLVKNIGT SKLIGQNGEE SEKEKEDTPE HDEEEEGKPA RVIKDEDKDS NLKNDEEGKN 300
301 SKSKDAQDND EEEEEGESDE EEEEGEEERR NIEESNNPSL RDVIVCCSEY SNIKELPHPI 360
361 DLTKKKCIFQ IINPVAPHHI GWAIGAFNSW SLPLISPPSV DAEDEVEEDK LRENVVDNVN 420
421 DTMDDDIGSD IIPIQIFTLP TQETDELTVI NSTVVCQKII DFYSKEFGSY PFTCYSMVFL 480
481 PTAPSKHMDF AALGICNTRL LYPLEVIDKA FSTTNELAWA LANQWSCVNI TPLDMNDYWC 540
541 CLGIAGYMVF QVTKKLMGNN TYKYQLKRNS EAIVEQDFEK PPIGSTFTGS SRPISWSSKD 600
601 LSFIQLKAPM ILHILDRRMT KTERSFGMSR VLPKIFLQAM SGDLPNNSLT SSHFQHVCER 660
661 VNKSKLENFF NEWVYGSGVP ILRVTQRFNR KRMVIELGIR QVQDEELGHE KVVGEEGFFK 720
721 SALDHLEHPD LNRTECFTGS MTIRIHEHDG TPYEHIVEIK DTFTKIDIQY NTKYRRLRKR 780
781 GGGANDENGV ENNNEEKPIV VDVNCLGNVY MSPEECSRFS LTEFNRTSES NELLKQNEAF 840
841 EWIRIDSDLE WICQMHINQP DYMFSSQLRQ DGDIEAQLEA IRYYEDVVVN GGVKSLVYSS 900
901 ILFRTAIDER YFFGIRLAAC EALSKYVYDP DFTGGVKHLI QIFQILFCLE DSNIPKSNNF 960
961 ENPKLYFLQC NIPKYLAKVK NENGKCPKLV KQFLLDILVY NENGENKYSD DAYVRSLIEN1020
1021 VVKVALNEYK DKAYMEKVKT QLLRYENLVN WLSSYESLIK TTIMYAKYKL HKVGAYDFTE1080
1081 LTGMIMHTLT LGINNGDISR ESFQNEFLMV LKIMLLEGGL KNKDALVLFT EILCFHEDSY1140
1141 IRDKSVDVLS ECVNLVVMDG SLDTISDDIK SSVQSVHNEV KNIKSEDDIE LFLSGHYVDD1200
1201 MKIKIEKIGR QNISGLIQIC RDMFKGYSPL KILLWDVLNL PVLSLYQRKQ IHDLVRVMYT1260
1261 LINSFVVRLE TPRERRLVAK MNSNEEGKLD IVIKRESILK VHIKKEVTST VEAPKKANKI1320
1321 KISLKGDKPV RKVEKQIVKP KVTSKQRKVK SHVNRMGSLP LRFVKIQQQP RVMVHLSSVP1380
1381 YSQFVQITKV TSRSFMVKIR TKNDAKN |
| PUBLICATION | TOPOLOGY | COCOMPLEXED PROTEINS | |
| View Details | Riffle et al. (2010) (Unpublished Data) |

|
TAF2 TAF8 |
| View Details | (MIPS) Mewes HW, et al. (2004) |
|
TAF1 TAF10 TAF11 TAF12 TAF13 TAF14 TAF2 TAF3 TAF5 TAF6 TAF7 TAF9 |
| View Details | Krogan NJ, et al. (2006) |
|
ESC1 ROX1 TAF2 TAF8 |
| View Details | Gavin AC, et al. (2006) |

|
TAF2 TAF8 |
| View Details | Gavin AC, et al. (2002) |

|
ADA2 ADH1 CLU1 FUN19 GCN5 HEM13 KAP114 KAP123 MED4 MES1 MNN4 MOT1 MYO1 NGG1 NOP1 PFK2 PSK1 PTC1 PUF6 RPB2 RPN9 RPO21 RPT5 RVB2 SGF29 SGF73 SIN3 SIN4 SPT15 SPT20 SPT7 SPT8 TAF1 TAF10 TAF11 TAF12 TAF13 TAF14 TAF2 TAF3 TAF4 TAF5 TAF6 TAF7 TAF8 TAF9 TBF1 TEF4 VAS1 VPS1 |
| View Details | Qiu et al. (2008) |
|
SGF73 SPT20 SPT7 SSL2 TAF1 TAF10 TAF11 TAF12 TAF13 TAF14 TAF2 TAF3 TAF4 TAF5 TAF6 TAF7 TAF9 TFA1 TFA2 TFB1 TFB5 TFC4 TFG1 TFG2 |
The following runs contain data for this protein:
| BAIT | COMMENTS | PUBLICATION | |
| View Run | sample cbf3 from feb 2005 | Sandall S, et all (2006) | |
| View Run | Sample 2- cpn2 from june 2005 | McCusker D, et al (2007) | |
| View Run | #32 Asynchronous Prep-No Phosphopeptide enrichment | Keck JM, et al. (2011) | |
| View Run | #29 Asynchronous Prep5-TiO2 Flowthrough | Keck JM, et al. (2011) |
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
New Feature: Upload Your Own Microscopy Data
| PROTEIN(S) | PUBLICATION | |
| View Data |
|
Huh WK, et al. (2003) |
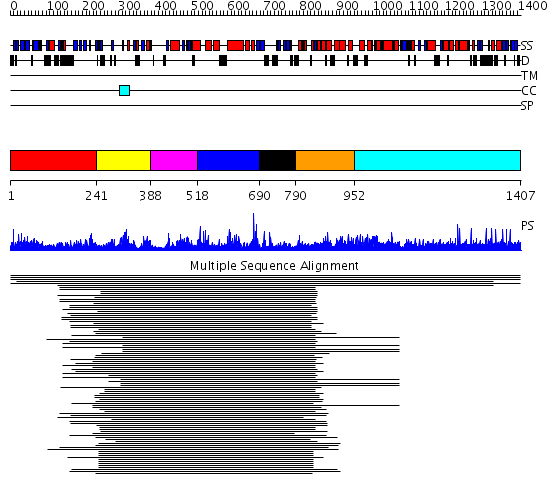
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..240] | 11.15 | Leukotriene A4 hydrolase C-terminal domain; Leukotriene A4 hydrolase N-terminal domain; Leukotriene A4 hydrolase catalytic domain | |
| 2 | View Details | [241..387] | 80.0 | Leukotriene A4 hydrolase C-terminal domain; Leukotriene A4 hydrolase N-terminal domain; Leukotriene A4 hydrolase catalytic domain | |
| 3 | View Details | [388..517] | 80.0 | Leukotriene A4 hydrolase C-terminal domain; Leukotriene A4 hydrolase N-terminal domain; Leukotriene A4 hydrolase catalytic domain | |
| 4 | View Details | [518..689] | 80.0 | Leukotriene A4 hydrolase C-terminal domain; Leukotriene A4 hydrolase N-terminal domain; Leukotriene A4 hydrolase catalytic domain | |
| 5 | View Details | [690..789] | 80.0 | Leukotriene A4 hydrolase C-terminal domain; Leukotriene A4 hydrolase N-terminal domain; Leukotriene A4 hydrolase catalytic domain | |
| 6 | View Details | [790..951] | 2.127996 | View MSA. No confident structure predictions are available. | |
| 7 | View Details | [952..1407] | N/A | No confident structure predictions are available. |
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.87 |
Source: Reynolds et al. (2008)