













| General Information: |
|
| Name(s) found: |
RFC2 /
YJR068W
[SGD]
|
| Description(s) found:
Found 33 descriptions. SHOW ALL |
|
| Organism: | Saccharomyces cerevisiae |
| Length: | 353 amino acids |
Gene Ontology: |
|
| Cellular Component: |
DNA replication factor C complex
[IDA |
| Biological Process: |
sister chromatid cohesion
[IPI cell cycle checkpoint [IMP leading strand elongation [IDA mismatch repair [TAS |
| Molecular Function: |
DNA clamp loader activity
[IDA purine nucleotide binding [ISS |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MFEGFGPNKK RKISKLAAEQ SLAQQPWVEK YRPKNLDEVT AQDHAVTVLK KTLKSANLPH 60
61 MLFYGPPGTG KTSTILALTK ELYGPDLMKS RILELNASDE RGISIVREKV KNFARLTVSK 120
121 PSKHDLENYP CPPYKIIILD EADSMTADAQ SALRRTMETY SGVTRFCLIC NYVTRIIDPL 180
181 ASRCSKFRFK ALDASNAIDR LRFISEQENV KCDDGVLERI LDISAGDLRR GITLLQSASK 240
241 GAQYLGDGKN ITSTQVEELA GVVPHDILIE IVEKVKSGDF DEIKKYVNTF MKSGWSAASV 300
301 VNQLHEYYIT NDNFDTNFKN QISWLLFTTD SRLNNGTNEH IQLLNLLVKI SQL |
| PUBLICATION | TOPOLOGY | COCOMPLEXED PROTEINS | |
| View Details | Riffle et al. (2010) (Unpublished Data) |
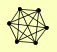
|
ELG1 RFC1 RFC2 RFC3 RFC4 RFC5 |
| View Details | (MIPS) Mewes HW, et al. (2004) |
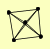
|
RFC1 RFC2 RFC3 RFC4 RFC5 |
| View Details | Krogan NJ, et al. (2006) |
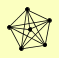
|
ELG1 RFC1 RFC2 RFC3 RFC4 RFC5 |
| View Details | Gavin AC, et al. (2006) |

|
RFC1 RFC2 |
| View Details | Gavin AC, et al. (2002) |

|
CTF18 CTF8 DPB2 ELG1 OLA1 POL2 RFC1 RFC2 RFC3 RFC4 RFC5 TOP2 VPS13 YMR181C |
| View Details | Ho Y, et al. (2002) |

|
ACH1 ACO1 ADE5,7 ADH2 AHA1 ARP2 ATP3 BRR2 CCT3 CLU1 CPA2 DUN1 ELG1 EXG1 FAA4 FET3 GUS1 HEF3 HSP104 IDH1 MDH2 NPA3 NTG1 PDI1 PGM2 PRB1 RAD24 RFC1 RFC2 RFC3 RFC4 RFC5 RNQ1 ROM2 RPC40 RPN5 RPT3 SAN1 SRP1 TCP1 TIF34 VAC8 YHR020W YIL108W YLR413W |
| View Details | Qiu et al. (2008) |
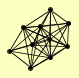
|
CDC45 DPB11 DPB2 MCM6 POL31 RFC1 RFC2 RFC3 RFC4 RFC5 |
The following runs contain data for this protein:
| BAIT | COMMENTS | PUBLICATION | |
| View Run | No Comments | Schneider, DA, et al. (2006) |
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
New Feature: Upload Your Own Microscopy Data
| PROTEIN(S) | PUBLICATION | |
| View Data |
|
Huh WK, et al. (2003) |
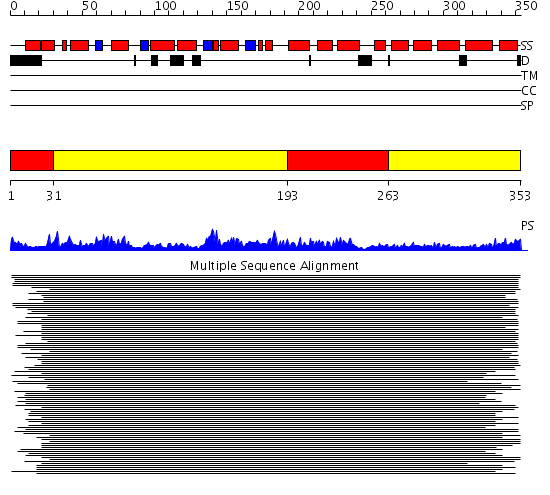

Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..30] [193..262] |
679.315398 | Replication factor C | |
| 2 | View Details | [31..192] [263..353] |
679.315398 | Replication factor C |
Functions predicted (by domain):
| # | Gene Ontology predictions | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 1 |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 2 | No functions predicted. |
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.99 |
Source: Reynolds et al. (2008)