













| General Information: |
|
| Name(s) found: |
CYS3 /
YAL012W
[SGD]
|
| Description(s) found:
Found 26 descriptions. SHOW ALL |
|
| Organism: | Saccharomyces cerevisiae |
| Length: | 394 amino acids |
Gene Ontology: |
|
| Cellular Component: |
cytoplasm
[IDA |
| Biological Process: |
sulfur amino acid metabolic process
[TAS response to drug [IMP transsulfuration [TAS cysteine metabolic process [TAS |
| Molecular Function: |
cystathionine gamma-lyase activity
[IDA |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MTLQESDKFA TKAIHAGEHV DVHGSVIEPI SLSTTFKQSS PANPIGTYEY SRSQNPNREN 60
61 LERAVAALEN AQYGLAFSSG SATTATILQS LPQGSHAVSI GDVYGGTHRY FTKVANAHGV 120
121 ETSFTNDLLN DLPQLIKENT KLVWIETPTN PTLKVTDIQK VADLIKKHAA GQDVILVVDN 180
181 TFLSPYISNP LNFGADIVVH SATKYINGHS DVVLGVLATN NKPLYERLQF LQNAIGAIPS 240
241 PFDAWLTHRG LKTLHLRVRQ AALSANKIAE FLAADKENVV AVNYPGLKTH PNYDVVLKQH 300
301 RDALGGGMIS FRIKGGAEAA SKFASSTRLF TLAESLGGIE SLLEVPAVMT HGGIPKEARE 360
361 ASGVFDDLVR ISVGIEDTDD LLEDIKQALK QATN |
| PUBLICATION | TOPOLOGY | COCOMPLEXED PROTEINS | |
| View Details | Krogan NJ, et al. (2006) |
|
CYS3 IPL1 RAX2 SER1 UBA1 YDR415C YER156C |
| View Details | Gavin AC, et al. (2006) |

|
CYS3 RNA1 |
| View Details | Gavin AC, et al. (2002) |
|
ADE4 CYS3 RNA1 |
| View Details | Ho Y, et al. (2002) |
|
ACS2 ADE3 ADE6 ADO1 ALA1 APE2 ARA1 ATG5 BAT1 CDC10 CLU1 CYS3 DED81 DPS1 ERG10 ERG20 FET3 GCY1 GPH1 GRS1 HOM2 HYP2 ILS1 IMD3 KRS1 LSP1 MDH3 OLA1 PAB1 PFK2 PGM2 PMI40 PRP6 SAC6 SCP160 SOD1 SSZ1 THR4 TRR1 UBA1 YDL124W |
The following runs contain data for this protein:
| BAIT | COMMENTS | PUBLICATION | |
| View Run | 3912: TAP-tagged, strain background includes hpc2(delta) | Green EM, et al (2005) | |
| View Run | #25 Asynchronous Prep3-TiO2 Flowthrough | Keck JM, et al. (2011) | |
| View Run | #24 Asynchronous Prep3-TiO2 Phosphopeptide enrichment | Keck JM, et al. (2011) | |
| View Run | #32 Asynchronous Prep-No Phosphopeptide enrichment | Keck JM, et al. (2011) | |
| View Run | #04 Alpha Factor Prep2 | Keck JM, et al. (2011) | |
| View Run | #14 Mitotic Prep2-TiO2 Flowthrough | Keck JM, et al. (2011) | |
| View Run | #12 Mitotic Prep1-TiO2 Flowthrough | Keck JM, et al. (2011) | |
| View Run | #02 Alpha Factor Prep1-TiO2 enriched, new search criteria | Keck JM, et al. (2011) | |
| View Run | #30 Asynchronous Prep6-TiO2 Phosphopeptide enrichment | Keck JM, et al. (2011) | |
| View Run | #11 Mitotic Prep1-TiO2 enriched, new search criteria | Keck JM, et al. (2011) |
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
| BAIT | PREY | HITS | PUBLICATION | |
| View Screen | 2 | Unpublished Fields Lab Data |
New Feature: Upload Your Own Microscopy Data
| PROTEIN(S) | PUBLICATION | |
| View Data |
|
Huh WK, et al. (2003) |
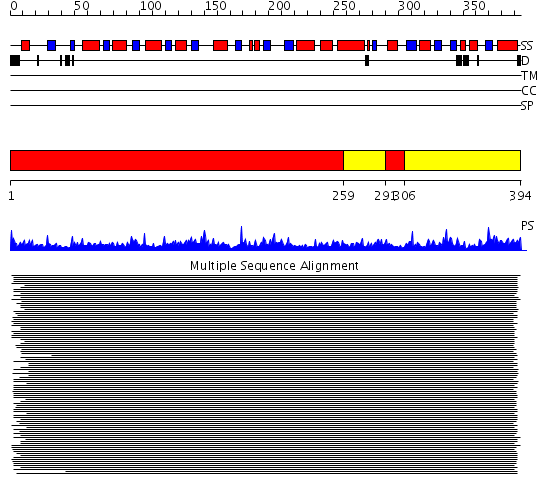
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..258] [291..305] |
896.228787 | Methionine gamma-lyase, MGL | |
| 2 | View Details | [259..290] [306..394] |
896.228787 | Methionine gamma-lyase, MGL |
Functions predicted (by domain):
| # | Gene Ontology predictions | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 1 |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 2 | No functions predicted. |
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.96 |
Source: Reynolds et al. (2008)