













| General Information: |
|
| Name(s) found: |
l(1)G0007-PB /
FBpp0073718
[FlyBase]
|
| Description(s) found:
Found 20 descriptions. SHOW ALL |
|
| Organism: | Drosophila melanogaster |
| Length: | 534 amino acids |
Gene Ontology: |
|
| Cellular Component: |
spliceosomal complex
[ISS]
|
| Biological Process: |
regulation of alternative nuclear mRNA splicing, via spliceosome
[IMP]
inter-male aggressive behavior [IMP] nuclear mRNA splicing, via spliceosome [ISS] |
| Molecular Function: |
nucleic acid binding
[IEA]
ATP binding [IEA] ATP-dependent helicase activity [ISS] ATP-dependent RNA helicase activity [ISS] |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MDSSKFATFF GNVPTFTIPG RTFPVDVMFS KNTCEDYVES AVKQALQVHL TPNEGDMLIF 60
61 MPGQEDIEVT CEVLEERLAE IDNAPALSIL PIYSQLPSDL QAKIFQKSSD GLRKCVVATN 120
121 IAETSLTVDG IIYVIDSGYC KLKVYNPRIG MDALQIYPIS QANANQRSGR AGRTGPGQAY 180
181 RLYTQRQYKD ELLALTVPEI QRTNLANTVL LLKSLGVVDL LQFHFMDPPP QDNILNSLYQ 240
241 LWILGALDHT GALTTLGRQM AEFPLDPPQC QMLIVACRMG CSAEVLIIVS MLSVPSIFYR 300
301 PKGREDEADG VREKFQRPES DHLTYLNVYQ QWRQNNYSST WCNEHFIHIK AMRKVREVRQ 360
361 QLKDIMTQQN LSVISCGIDW DIVRKCICSA YFYQAARLKG IGEYVNLRTG MPCHLHPTSA 420
421 LYGLGTTPDY VVYHELIMTA KEYMQCATAV DGYWLAELGP MFFSVKESGR SGREKKKQAA 480
481 EHLKEMEEQM LKAQHEMEER KQQAAEREEQ LATKQEIATP GNATPRRTPA RIGL |
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
New Feature: Upload Your Own Microscopy Data
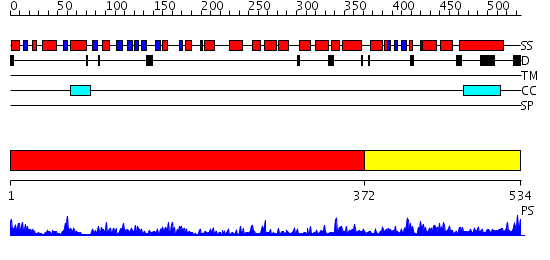
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..371] | 13.522879 | Structure of the RecQ Catalytic Core | |
| 2 | View Details | [372..534] | 20.619789 | No description for PF07717.7 was found. No confident structure predictions are available. |
Functions predicted (by domain):
| # | Gene Ontology predictions | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 1 |
|
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 2 | No functions predicted. |
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.86 |
Source: Reynolds et al. (2008)