













| General Information: |
|
| Name(s) found: |
AIR1 /
YIL079C
[SGD]
|
| Description(s) found:
Found 28 descriptions. SHOW ALL |
|
| Organism: | Saccharomyces cerevisiae |
| Length: | 360 amino acids |
Gene Ontology: |
|
| Cellular Component: |
nucleus
[IPI nucleolus [IDA TRAMP complex [IDA |
| Biological Process: |
mRNA export from nucleus
[IMP rRNA catabolic process [IGI snRNA catabolic process [IGI snoRNA catabolic process [IGI tRNA catabolic process [IDA ncRNA polyadenylation involved in polyadenylation-dependent ncRNA catabolic process [IDA |
| Molecular Function: |
polynucleotide adenylyltransferase activity
[IDA |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MSTLLSEVES IDTLPYVKDT TPTGSDSSSF NKLLAPSIED VDANPEELRT LRGQGRYFGI 60
61 TDYDSNGAIM EAEPKCNNCS QRGHLKRNCP HVICTYCGFM DDHYSQHCPK AIICTNCNAN 120
121 GHYKSQCPHK WKKVFCTLCN SKRHSRERCP SIWRSYLLKT KDANQGDFDF QTVFCYNCGN 180
181 AGHFGDDCAE RRSSRVPNTD GSAFCGDNLA TKFKQHYFNQ LKDYKREASQ RQHFDNEHEF 240
241 NLLDYEYNDD AYDLPGSRTY RDKMKWKGKV QSTRNKNSSN NRYESSNNRK KKSPFSAQNY 300
301 KVTKNKRVQT HPLDFPRSSQ NNRTNDYSSQ FSYNRDDFPK GPKNKRGRSS SNKSQRNGRY 360
361 |
| PUBLICATION | TOPOLOGY | COCOMPLEXED PROTEINS | |
| View Details | Riffle et al. (2010) (Unpublished Data) |
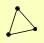
|
AIR1 MTR4 PAP2 |
| View Details | Krogan NJ, et al. (2006) |
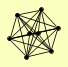
|
AIR1 AIR2 MTR4 PAP2 PSO2 RPL18A, RPL18B STE4 TRF5 |
| View Details | Ho Y, et al. (2002) |
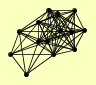
|
AIR1 AIR2 DED1 HRB1 IMD1 IMD3 IMD4 MTR4 NAP1 NOP53 NOP56 NPL3 PAP2 PSE1 |
| View Details | Qiu et al. (2008) |
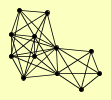
|
AIR1 AIR2 DBP7 DRS1 MTR4 NOG2 NOP1 NOP2 PAP2 RNT1 RRP3 TRF5 |
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
| BAIT | PREY | HITS | PUBLICATION | |
| View Screen | 2 | Unpublished Fields Lab Data | ||
| View Screen | 2 | Unpublished Fields Lab Data | ||
| View Screen | 2 | Drees BL, et al. (2001) |
New Feature: Upload Your Own Microscopy Data
| PROTEIN(S) | PUBLICATION | |
| View Data |
|
Huh WK, et al. (2003) |
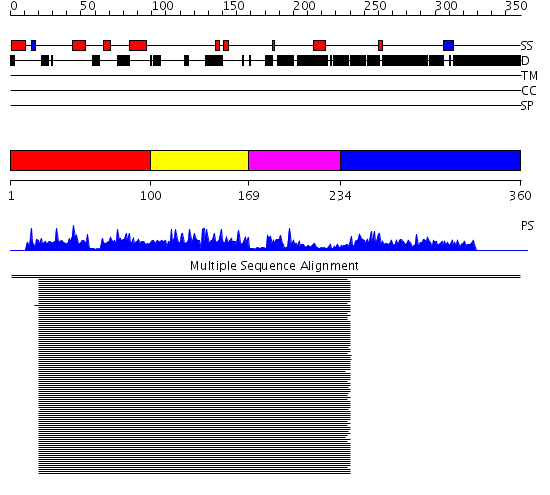
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..99] | 35.522879 | HIV capsid protein, dimerisation domain | |
| 2 | View Details | [100..168] | 24.69897 | HIV nucleocapsid | |
| 3 | View Details | [169..233] | N/A | No confident structure predictions are available. | |
| 4 | View Details | [234..360] | 25.029993 | View MSA. No confident structure predictions are available. |
Functions predicted (by domain):
| # | Gene Ontology predictions | ||||||||||||||||||||||||
| 1 |
|
||||||||||||||||||||||||
| 2 | No functions predicted. | ||||||||||||||||||||||||
| 3 | No functions predicted. | ||||||||||||||||||||||||
| 4 | No functions predicted. |
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.99 |
Source: Reynolds et al. (2008)