













| General Information: |
|
| Name(s) found: |
CAMK1_ARATH
[Swiss-Prot]
|
| Description(s) found:
Found 15 descriptions. SHOW ALL |
|
| Organism: | Arabidopsis thaliana |
| Length: | 576 amino acids |
Gene Ontology: |
|
| Cellular Component: |
plasma membrane
[IDA]
|
| Biological Process: |
protein amino acid phosphorylation
[IEA]
|
| Molecular Function: |
calcium-dependent protein serine/threonine phosphatase activity
[ISS]
calcium ion binding [ISS] calmodulin-dependent protein kinase activity [IDA] kinase activity [ISS] |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MGICHGKPVE QQSKSLPVSG ETNEAPTNSQ PPAKSSGFPF YSPSPVPSLF KSSPSVSSSV 60
61 SSTPLRIFKR PFPPPSPAKH IRAFLARRYG SVKPNEVSIP EGKECEIGLD KSFGFSKQFA 120
121 SHYEIDGEVG RGHFGYTCSA KGKKGSLKGQ EVAVKVIPKS KMTTAIAIED VSREVKMLRA 180
181 LTGHKNLVQF YDAFEDDENV YIVMELCKGG ELLDKILQRG GKYSEDDAKK VMVQILSVVA 240
241 YCHLQGVVHR DLKPENFLFS TKDETSPLKA IDFGLSDYVK PDERLNDIVG SAYYVAPEVL 300
301 HRTYGTEADM WSIGVIAYIL LCGSRPFWAR TESGIFRAVL KAEPNFEEAP WPSLSPEAVD 360
361 FVKRLLNKDY RKRLTAAQAL CHPWLVGSHE LKIPSDMIIY KLVKVYIMST SLRKSALAAL 420
421 AKTLTVPQLA YLREQFTLLG PSKNGYISMQ NYKTAILKSS TDAMKDSRVF DFVHMISCLQ 480
481 YKKLDFEEFC ASALSVYQLE AMETWEQHAR RAYELFEKDG NRPIMIEELA SELGLGPSVP 540
541 VHVVLQDWIR HSDGKLSFLG FVRLLHGVSS RTLQKA |
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
New Feature: Upload Your Own Microscopy Data
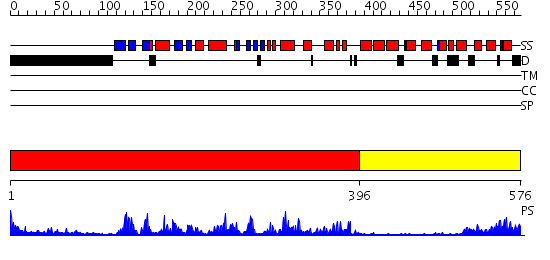
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..395] | 54.045757 | Abl tyrosine kinase, SH3 domain; Abl tyrosine kinase; Abelsone tyrosine kinase (abl) | |
| 2 | View Details | [396..576] | 21.30103 | Ca2+-regulatory region (CLD) from soybean calcium-dependent protein kinase-alpha (CDPK) in the presence of Ca2+ and the junction domain (JD) |
Functions predicted (by domain):
| # | Gene Ontology predictions | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 1 |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 2 |
|
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.89 |
Source: Reynolds et al. (2008)