













| General Information: |
|
| Name(s) found: |
LCB2 /
YDR062W
[SGD]
|
| Description(s) found:
Found 32 descriptions. SHOW ALL |
|
| Organism: | Saccharomyces cerevisiae |
| Length: | 561 amino acids |
Gene Ontology: |
|
| Cellular Component: |
membrane fraction
[IDA microsome [IPI serine C-palmitoyltransferase complex [IMP |
| Biological Process: |
sphingolipid biosynthetic process
[TAS |
| Molecular Function: |
serine C-palmitoyltransferase activity
[IDA |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MSTPANYTRV PLCEPEELPD DIQKENEYGT LDSPGHLYQV KSRHGKPLPE PVVDTPPYYI 60
61 SLLTYLNYLI LIILGHVHDF LGMTFQKNKH LDLLEHDGLA PWFSNFESFY VRRIKMRIDD 120
121 CFSRPTTGVP GRFIRCIDRI SHNINEYFTY SGAVYPCMNL SSYNYLGFAQ SKGQCTDAAL 180
181 ESVDKYSIQS GGPRAQIGTT DLHIKAEKLV ARFIGKEDAL VFSMGYGTNA NLFNAFLDKK 240
241 CLVISDELNH TSIRTGVRLS GAAVRTFKHG DMVGLEKLIR EQIVLGQPKT NRPWKKILIC 300
301 AEGLFSMEGT LCNLPKLVEL KKKYKCYLFI DEAHSIGAMG PTGRGVCEIF GVDPKDVDIL 360
361 MGTFTKSFGA AGGYIAADQW IIDRLRLDLT TVSYSESMPA PVLAQTISSL QTISGEICPG 420
421 QGTERLQRIA FNSRYLRLAL QRLGFIVYGV ADSPVIPLLL YCPSKMPAFS RMMLQRRIAV 480
481 VVVAYPATPL IESRVRFCMS ASLTKEDIDY LLRHVSEVGD KLNLKSNSGK SSYDGKRQRW 540
541 DIEEVIRRTP EDCKDDKYFV N |
| PUBLICATION | TOPOLOGY | COCOMPLEXED PROTEINS | |
| View Details | Riffle et al. (2010) (Unpublished Data) |

|
LCB1 LCB2 |
| View Details | Gavin AC, et al. (2006) |

|
LCB1 LCB2 |
| View Details | Gavin AC, et al. (2002) |

|
BMH1 BMH2 CLU1 CRM1 ECM29 FKS1 GCN1 GEA2 GFA1 HRP1 KAP104 KAP123 LCB1 LCB2 LYS12 MIR1 NAB2 SAM1 SAM2 YHR020W |
| View Details | Qiu et al. (2008) |

|
LCB1 LCB2 PIS1 SUR4 |
The following runs contain data for this protein:
| BAIT | COMMENTS | PUBLICATION | |
| View Run | #03 Alpha Factor Prep1-TiO2 Flowthrough | Keck JM, et al. (2011) |
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
New Feature: Upload Your Own Microscopy Data
| PROTEIN(S) | PUBLICATION | |
| View Data |
|
Huh WK, et al. (2003) |
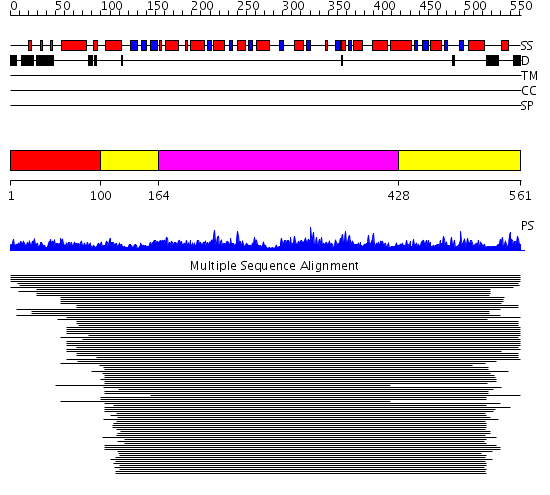
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..99] | N/A | No confident structure predictions are available. | |
| 2 | View Details | [100..163] [428..561] |
499.325697 | 2-amino-3-ketobutyrate CoA ligase | |
| 3 | View Details | [164..427] | 499.325697 | 2-amino-3-ketobutyrate CoA ligase |
Functions predicted (by domain):
| # | Gene Ontology predictions | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 1 | No functions predicted. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 2 |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 3 | No functions predicted. |
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.48 |
Source: Reynolds et al. (2008)