













| General Information: |
|
| Name(s) found: |
ATG7 /
YHR171W
[SGD]
|
| Description(s) found:
Found 26 descriptions. SHOW ALL |
|
| Organism: | Saccharomyces cerevisiae |
| Length: | 630 amino acids |
Gene Ontology: |
|
| Cellular Component: |
cytoplasm
[IDA mitochondrion [IDA membrane [IDA cytosol [IDA |
| Biological Process: |
autophagy
[IMP protein modification process [IMP C-terminal protein lipidation [IDA protein targeting to vacuole [IMP CVT pathway [IMP |
| Molecular Function: |
APG8 activating enzyme activity
[IMP APG12 activating enzyme activity [IMP |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MSSERVLSYA PAFKSFLDTS FFQELSRLKL DVLKLDSTCQ PLTVNLDLHN IPKSADQVPL 60
61 FLTNRSFEKH NNKRTNEVPL QGSIFNFNVL DEFKNLDKQL FLHQRALECW EDGIKDINKC 120
121 VSFVIISFAD LKKYRFYYWL GVPCFQRPSS TVLHVRPEPS LKGLFSKCQK WFDVNYSKWV 180
181 CILDADDEIV NYDKCIIRKT KVLAIRDTST MENVPSALTK NFLSVLQYDV PDLIDFKLLI 240
241 IRQNEGSFAL NATFASIDPQ SSSSNPDMKV SGWERNVQGK LAPRVVDLSS LLDPLKIADQ 300
301 SVDLNLKLMK WRILPDLNLD IIKNTKVLLL GAGTLGCYVS RALIAWGVRK ITFVDNGTVS 360
361 YSNPVRQALY NFEDCGKPKA ELAAASLKRI FPLMDATGVK LSIPMIGHKL VNEEAQHKDF 420
421 DRLRALIKEH DIIFLLVDSR ESRWLPSLLS NIENKTVINA ALGFDSYLVM RHGNRDEQSS 480
481 KQLGCYFCHD VVAPTDSLTD RTLDQMCTVT RPGVAMMASS LAVELMTSLL QTKYSGSETT 540
541 VLGDIPHQIR GFLHNFSILK LETPAYEHCP ACSPKVIEAF TDLGWEFVKK ALEHPLYLEE 600
601 ISGLSVIKQE VERLGNDVFE WEDDESDEIA |
| PUBLICATION | TOPOLOGY | COCOMPLEXED PROTEINS | |
| View Details | Ho Y, et al. (2002) |

|
ATG7 NOP2 TIF6 ULS1 |
The following runs contain data for this protein:
| BAIT | COMMENTS | PUBLICATION | |
| View Run | sample cbf3 from feb 2005 | Sandall S, et all (2006) |
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
New Feature: Upload Your Own Microscopy Data
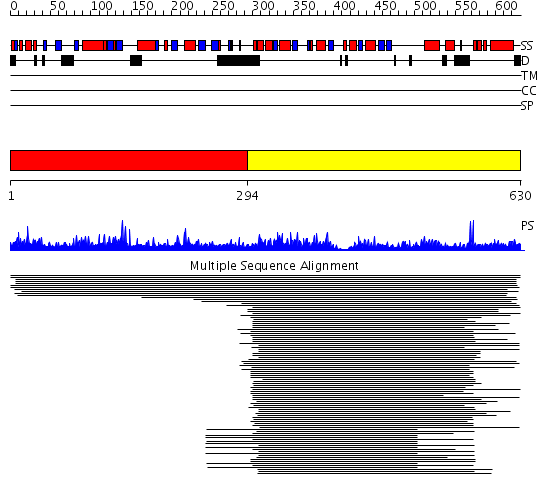
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..293] | N/A | No confident structure predictions are available. | |
| 2 | View Details | [294..630] | 166.30103 | Molybdenum cofactor biosynthesis protein MoeB | |
| 3 | View Details | [300..630] | 69.39794 | Molybdenum cofactor biosynthesis protein MoeB |
Functions predicted (by domain):
| # | Gene Ontology predictions | ||||||||||||||||||||||||||||||||||||
| 1 | No functions predicted. | ||||||||||||||||||||||||||||||||||||
| 2 | No functions predicted. | ||||||||||||||||||||||||||||||||||||
| 3 |
|
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.96 |
Source: Reynolds et al. (2008)