













| General Information: |
|
| Name(s) found: |
PTN2A_ARATH
[Swiss-Prot]
|
| Description(s) found:
Found 13 descriptions. SHOW ALL |
|
| Organism: | Arabidopsis thaliana |
| Length: | 611 amino acids |
Gene Ontology: |
|
| Cellular Component: | NONE FOUND |
| Biological Process: |
dephosphorylation
[IEA]
|
| Molecular Function: |
protein tyrosine phosphatase activity
[IEA]
phosphatase activity [IEA] |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MSSESPNLPA AAGTVPDNHP PPPPVVTAAE AGSDDSPKGV ASKLSAAGIS NWAKNLKVPQ 60
61 PFASTQNDSG VENTEKSAFA KFTSGLGIRL SPKSPQTNDT TTEGTSSATE SSFIGTITKG 120
121 LVDTSKNAVK AVQVKARHAV SQNKRRYQEG GFDLDLTYIT ENIIAMGFPA GDMSSGFFGY 180
181 VEGFYRNQME EVINFLETQH KGKYKVYNLC SERLYDVSLF EGKVASFPFD DHNCPPIHLV 240
241 TSFCQSAYSW LKEDIENVVV VHCKAGMART GLMICSLLLY LKFFPTAEEC MDFYNQKRCV 300
301 DGKGLVLPSQ IRYVKYFERI LTYFNGENQP GRRCMLRGFR LHRCPYWIRP SITISDHNGV 360
361 LFTTKKHPRT KDLSPEDFWF SAPKKGVMVF ALPGEPGLTE LAGDFKIQFH DRQGDFYCWL 420
421 NTTMMENRVI LKTSELDGFD KRKLPSPGFM VEVVLADINA TIPTNPSSET ASKTPEETSA 480
481 ANSSPVDGSA SVPGPDKETE NPDKDDVFSD NEGDSTGPTK TTSSASSQTP EAKKSADETA 540
541 VLTKATEKVS ISGNKGSSQP VQGVTVSKGE ATEKPSGAGV NASSSSESEF KVMAADASVF 600
601 SFGDEDDFES D |
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
New Feature: Upload Your Own Microscopy Data
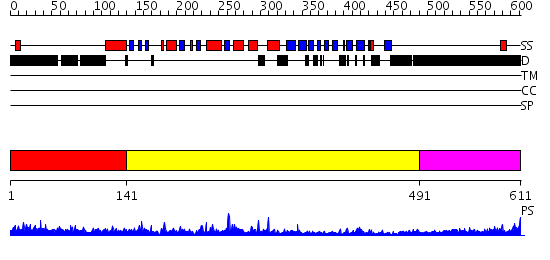
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..140] | 56.69897 | Proline directed phosphatase CDC14b2 | |
| 2 | View Details | [141..490] | 68.09691 | Pten tumor suppressor (Phoshphoinositide phosphatase), C-terminal domain; Phoshphoinositide phosphatase Pten (Pten tumor suppressor), N-terminal domain | |
| 3 | View Details | [491..611] | 2.201996 | View MSA. No confident structure predictions are available. |
Functions predicted (by domain):
| # | Gene Ontology predictions | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 1 |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 2 |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 3 | No functions predicted. |
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.98 |
Source: Reynolds et al. (2008)