













| General Information: |
|
| Name(s) found: |
rpn502 /
SPAPB8E5.02c
[Sanger Pombe]
rpn501 / SPAC1420.03 [Sanger Pombe] |
| Description(s) found:
Found 29 descriptions. SHOW ALL |
|
| Organism: | Schizosaccharomyces pombe |
| Length: | 443 amino acids |
Gene Ontology: |
|
| Cellular Component: |
nuclear envelope
[IDA nucleus [IDA cytosol [IDA proteasome complex [ISS nuclear periphery [IDA |
| Biological Process: |
proteasomal ubiquitin-dependent protein catabolic process
[IGI proteasome localization [IGI regulation of cell cycle [RCA proteasome assembly [IGI mitotic cell cycle [IC] |
| Molecular Function: |
protein binding
[IPI |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MEQKPEVDYS EKFAELQKSL NNLNTIDIDA NLEKLLIFEK QVRQASDTST NTKVLIYIAD 60
61 LLFRAGDFQG LNEQLVSLFK KHGQLKQSMT SLVQHVMTYL PGIDDLKTKI NLIETLRTIT 120
121 DGKIYVEVER ARLTQLLSQI KEEQGDIKSA QEILCNEPVE TYGSFDLKEK VAFILDQVRL 180
181 FLLRSDYYMA STYTKKINVK FFEKEDVQSL KLKYYEQKIR IGLHDDAYLD VCKYYRAVYD 240
241 TAVVQEDPEK WKEILENVVC FALLTPYDNE QADLLHRINA DHKLNSLPLL QQLVKCFIVN 300
301 ELMRWPKIAE IYGSALRSNP VFAENDEKGE KRWSELRKRV IEHNIRVVAN YYSRIHCSRL 360
361 GVLLDMSPSE TEQFLCDLIA KHHFYAKIDR PAQVISFKKS QNVQEQLNEW GSNITELLGK 420
421 LEKVRQLIIK EEMMNSIQQA VAK |
SHOWING SINGLE HITS. [ Hide Single Hits ]
New Feature: Upload Your Own Microscopy Data
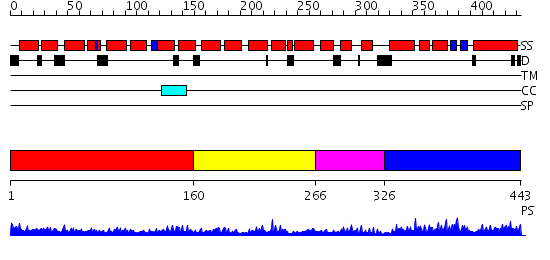
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..159] | 7.045757 | Crystal structure of an 8 repeat consensus TPR superhelix (trigonal crystal form) | |
| 2 | View Details | [160..265] | 5.0 | Designed protein cTPR3 | |
| 3 | View Details | [266..325] | N/A | No confident structure predictions are available. | |
| 4 | View Details | [326..443] | 12.09691 | Solution structure of the PCI domain |
Functions predicted (by domain):
| # | Gene Ontology predictions | ||||||||||||||||||||||||||||||||||||||||||||||||
| 1 | No functions predicted. | ||||||||||||||||||||||||||||||||||||||||||||||||
| 2 | No functions predicted. | ||||||||||||||||||||||||||||||||||||||||||||||||
| 3 | No functions predicted. | ||||||||||||||||||||||||||||||||||||||||||||||||
| 4 |
|
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.99 |
Source: Reynolds et al. (2008)