













| General Information: |
|
| Name(s) found: |
clr4 /
SPBC428.08c
[Sanger Pombe]
|
| Description(s) found:
SHOW ONLY BEST |
|
| Organism: | Schizosaccharomyces pombe |
| Length: | 490 amino acids |
Gene Ontology: |
|
| Cellular Component: |
mating-type region heterochromatin
[NAS nucleus [IDA nuclear telomeric heterochromatin [NAS CLRC ubiquitin ligase complex [IDA nuclear centromeric heterochromatin [TAS spindle pole body [IDA |
| Biological Process: |
regulation of production of siRNA involved in RNA interference
[IMP chromatin assembly or disassembly [IEA] donor selection [IMP chromatin silencing at telomere [TAS meiotic telomere clustering [IMP dynein-driven meiotic oscillatory nuclear movement [IGI chromatin remodeling [RCA chromatin silencing at centromere [IMP chromatin silencing at rDNA [IMP histone methylation [IDA regulation of Ran protein signal transduction [TAS chromatin silencing at silent mating-type cassette [IMP cellular protein localization [IGI attachment of spindle microtubules to kinetochore involved in mitotic sister chromatid segregation [IMP sphingolipid biosynthetic process [IMP |
| Molecular Function: |
zinc ion binding
[IEA]
protein binding [IPI histone methyltransferase activity (H3-K9 specific) [IDA chromatin binding [IEA] |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MSPKQEEYEV ERIVDEKLDR NGAVKLYRIR WLNYSSRSDT WEPPENLSGC SAVLAEWKRR 60
61 KRRLKGSNSD SDSPHHASNP HPNSRQKHQH QTSKSVPRSQ RFSRELNVKK ENKKVFSSQT 120
121 TKRQSRKQST ALTTNDTSII LDDSLHTNSK KLGKTRNEVK EESQKRELVS NSIKEATSPK 180
181 TSSILTKPRN PSKLDSYTHL SFYEKRELFR KKLREIEGPE VTLVNEVDDE PCPSLDFQFI 240
241 SQYRLTQGVI PPDPNFQSGC NCSSLGGCDL NNPSRCECLD DLDEPTHFAY DAQGRVRADT 300
301 GAVIYECNSF CSCSMECPNR VVQRGRTLPL EIFKTKEKGW GVRSLRFAPA GTFITCYLGE 360
361 VITSAEAAKR DKNYDDDGIT YLFDLDMFDD ASEYTVDAQN YGDVSRFFNH SCSPNIAIYS 420
421 AVRNHGFRTI YDLAFFAIKD IQPLEELTFD YAGAKDFSPV QSQKSQQNRI SKLRRQCKCG 480
481 SANCRGWLFG |
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
New Feature: Upload Your Own Microscopy Data
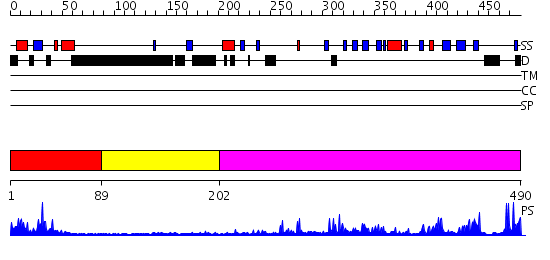
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..88] | 8.09691 | Histone methyltransferase clr4, chromo domain | |
| 2 | View Details | [89..201] | N/A | No confident structure predictions are available. | |
| 3 | View Details | [202..490] | 62.522879 | SET domain of Clr4 |
Functions predicted (by domain):
| # | Gene Ontology predictions | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 1 |
|
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 2 | No functions predicted. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 3 |
|
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.99 |
Source: Reynolds et al. (2008)