













| General Information: |
|
| Name(s) found: |
mutY /
EG10627
[EcoGene]
|
| Description(s) found:
Found 5 descriptions. SHOW ALL |
|
| Organism: | Escherichia coli str. K-12 substr. MG1655 |
| Length: | 350 amino acids |
Gene Ontology: |
|
| Cellular Component: |
intracellular
[IEA]
|
| Biological Process: |
DNA repair
[IEA]
base-excision repair [IEA] metabolic process [IEA] response to DNA damage stimulus [IEA] |
| Molecular Function: |
DNA binding
[IEA]
metal ion binding [IEA] iron ion binding [IEA] catalytic activity [IEA] endonuclease activity [IEA] DNA N-glycosylase activity [IEA] 4 iron, 4 sulfur cluster binding [IEA] iron-sulfur cluster binding [IEA] hydrolase activity, acting on glycosyl bonds [IEA] hydrolase activity [IEA] |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MQASQFSAQV LDWYDKYGRK TLPWQIDKTP YKVWLSEVML QQTQVATVIP YFERFMARFP 60
61 TVTDLANAPL DEVLHLWTGL GYYARARNLH KAAQQVATLH GGKFPETFEE VAALPGVGRS 120
121 TAGAILSLSL GKHFPILDGN VKRVLARCYA VSGWPGKKEV ENKLWSLSEQ VTPAVGVERF 180
181 NQAMMDLGAM ICTRSKPKCS LCPLQNGCIA AANNSWALYP GKKPKQTLPE RTGYFLLLQH 240
241 EDEVLLAQRP PSGLWGGLYC FPQFADEESL RQWLAQRQIA ADNLTQLTAF RHTFSHFHLD 300
301 IVPMWLPVSS FTGCMDEGNA LWYNLAQPPS VGLAAPVERL LQQLRTGAPV |
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
New Feature: Upload Your Own Microscopy Data
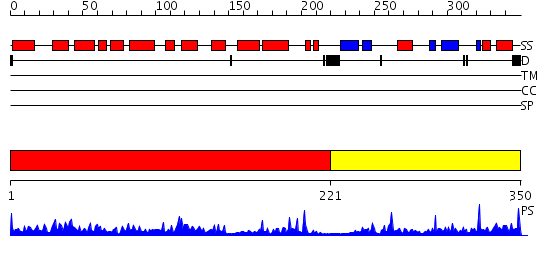
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..220] | 59.39794 | Catalytic domain of MutY | |
| 2 | View Details | [221..350] | 15.69897 | NMR STUDY OF MUTT ENZYME, A NUCLEOSIDE TRIPHOSPHATE PYROPHOSPHOHYDROLASE |
Functions predicted (by domain):
| # | Gene Ontology predictions | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 1 |
|
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 2 |
|
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.97 |
Source: Reynolds et al. (2008)