GO TERM SUMMARY
|
| Name: |
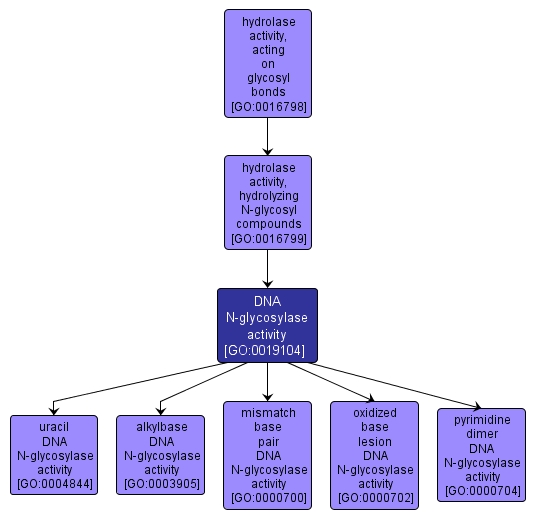
DNA N-glycosylase activity |
| Acc: |
GO:0019104 |
| Aspect: |
Molecular Function |
| Desc: |
Catalysis of the removal of damaged bases by cleaving the N-C1' glycosidic bond between the target damaged DNA base and the deoxyribose sugar. The reaction releases a free base and leaves an apurinic/apyrimidinic (AP) site. |
Synonyms:
- endonuclease VIII activity
- DNA glycosylase activity
- GO:0008578
|
|

|
INTERACTIVE GO GRAPH
|