













| General Information: |
|
| Name(s) found: |
DRP1A_ARATH
[Swiss-Prot]
|
| Description(s) found:
Found 15 descriptions. SHOW ALL |
|
| Organism: | Arabidopsis thaliana |
| Length: | 610 amino acids |
Gene Ontology: |
|
| Cellular Component: |
vacuole
[IDA]
cell plate [IDA] plasma membrane [IDA] microtubule [IDA] chloroplast thylakoid membrane [IDA] |
| Biological Process: |
embryonic development ending in seed dormancy
[IGI]
synaptic vesicle endocytosis [IDA] xylem and phloem pattern formation [IMP] cytokinesis by cell plate formation [IGI] trichome branching [IMP] cell plate formation involved in plant-type cell wall biogenesis [IGI] |
| Molecular Function: |
GTP binding
[ISS]
protein binding [IPI] GTPase activity [IDA] clathrin binding [IDA] |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MENLISLVNK IQRACTALGD HGDSSALPTL WDSLPAIAVV GGQSSGKSSV LESIVGKDFL 60
61 PRGSGIVTRR PLVLQLQKID DGTREYAEFL HLPRKKFTDF AAVRKEIQDE TDRETGRSKA 120
121 ISSVPIHLSI YSPNVVNLTL IDLPGLTKVA VDGQSDSIVK DIENMVRSYI EKPNCIILAI 180
181 SPANQDLATS DAIKISREVD PSGDRTFGVL TKIDLMDKGT DAVEILEGRS FKLKYPWVGV 240
241 VNRSQADINK NVDMIAARKR EREYFSNTTE YRHLANKMGS EHLAKMLSKH LERVIKSRIP 300
301 GIQSLINKTV LELETELSRL GKPIAADAGG KLYSIMEICR LFDQIFKEHL DGVRAGGEKV 360
361 YNVFDNQLPA ALKRLQFDKQ LAMDNIRKLV TEADGYQPHL IAPEQGYRRL IESSIVSIRG 420
421 PAEASVDTVH AILKDLVHKS VNETVELKQY PALRVEVTNA AIESLDKMRE GSKKATLQLV 480
481 DMECSYLTVD FFRKLPQDVE KGGNPTHSIF DRYNDSYLRR IGSNVLSYVN MVCAGLRNSI 540
541 PKSIVYCQVR EAKRSLLDHF FAELGTMDMK RLSSLLNEDP AIMERRSAIS KRLELYRAAQ 600
601 SEIDAVAWSK |
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
New Feature: Upload Your Own Microscopy Data
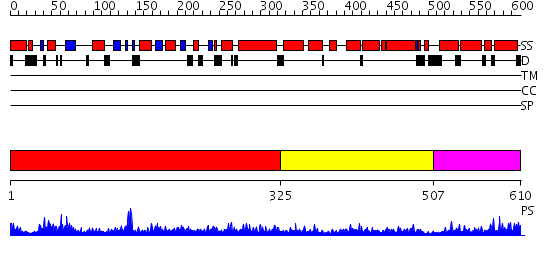
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..324] | 102.0 | Structure of the nucleotide-free myosin II motor domain from Dictyostelium discoideum fused to the GTPase domain of dynamin 1 from Rattus norvegicus | |
| 2 | View Details | [325..506] | 32.376751 | No description for PF01031.11 was found. No confident structure predictions are available. | |
| 3 | View Details | [507..610] | 39.958607 | No description for PF02212.9 was found. No confident structure predictions are available. |
Functions predicted (by domain):
| # | Gene Ontology predictions | ||||||||||||||||||||||||||||||
| 1 |
|
||||||||||||||||||||||||||||||
| 2 | No functions predicted. | ||||||||||||||||||||||||||||||
| 3 | No functions predicted. |
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.99 |
Source: Reynolds et al. (2008)