













| General Information: |
|
| Name(s) found: |
sum3 /
SPCC1795.11
[Sanger Pombe]
|
| Description(s) found:
Found 25 descriptions. SHOW ALL |
|
| Organism: | Schizosaccharomyces pombe |
| Length: | 636 amino acids |
Gene Ontology: |
|
| Cellular Component: |
cytoplasm
[NAS cytosol [IDA |
| Biological Process: |
regulation of translation
[IMP positive regulation of translational initiation [IC] DNA replication checkpoint [IGI G2/M transition of mitotic cell cycle [IMP response to osmotic stress [IMP gene silencing by RNA [IDA regulation of conjugation with cellular fusion [IGI |
| Molecular Function: |
translation initiation factor activity
[IEA]
ATP binding [IEA] protein binding [IPI RNA binding [IEA] ATP-dependent RNA helicase activity [TAS |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MSDNVQQQVD SVGSVTEKLQ KTNISRPRKY IPPFARDKPS AGAAPAVGDD ESVSSRGSSR 60
61 SQTPSEFSSN YGGRREYNRG GHYGGGEGRQ NNYRGGREGG YSNGGGYRNN RGFGQWRDGQ 120
121 HVIGARNTLL ERQLFGAVAD GTKVSTGINF EKYDDIPVEV SGGDIEPVNE FTSPPLNSHL 180
181 LQNIKLSGYT QPTPVQKNSI PIVTSGRDLM ACAQTGSGKT AGFLFPILSL AFDKGPAAVP 240
241 VDQDAGMGYR PRKAYPTTLI LAPTRELVCQ IHEESRKFCY RSWVRPCAVY GGADIRAQIR 300
301 QIDQGCDLLS ATPGRLVDLI DRGRISLANI KFLVLDEADR MLDMGFEPQI RHIVEGADMT 360
361 SVEERQTLMF SATFPRDIQL LARDFLKDYV FLSVGRVGST SENITQKVVH VEDSEKRSYL 420
421 LDILHTLPPE GLTLIFVETK RMADTLTDYL LNSNFPATSI HGDRTQRERE RALELFRSGR 480
481 TSIMVATAVA SRGLDIPNVT HVINYDLPTD IDDYVHRIGR TGRAGNTGQA VAFFNRNNKG 540
541 IAKELIELLQ EANQECPSFL IAMARESSFG GNGRGGRYSG RGGRGGNAYG ARDFRRPTNS 600
601 SSGYSSGPSY SGYGGFESRT PHHGNTYNSG SAQSWW |
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
New Feature: Upload Your Own Microscopy Data
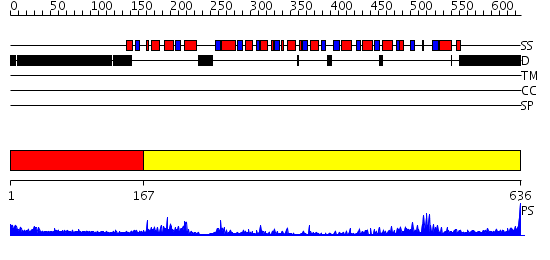
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..166] | 1.680995 | View MSA. No confident structure predictions are available. | |
| 2 | View Details | [167..636] | 96.045757 | No description for 2i4iA was found. |
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.99 |
Source: Reynolds et al. (2008)