













| General Information: |
|
| Name(s) found: |
FBpp0306674
[FlyBase]
Nrx-1-PA / FBpp0083649 [FlyBase] |
| Description(s) found:
SHOW ONLY BEST |
|
| Organism: | Drosophila melanogaster |
| Length: | 1837 amino acids |
Gene Ontology: |
|
| Cellular Component: |
plasma membrane
[NAS]
|
| Biological Process: |
synapse organization
[IDA]
synaptic transmission [IDA] |
| Molecular Function: | NONE FOUND |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MKAPHSATYQ DNYADAAMTA RTRPSMDMDQ QRNRNQAELR LLPAQRTSTS AFESPDLRFN 60
61 KARRRRRSEQ VPPVVVEYRR SYRVLIVSAL LVSLAASFVT LSAGFQLDGS QNSFYTFRKW 120
121 YTGLNGTLEL EFKTEQPNGL VLYTDDGGTY DFFELKLVEG ALRLRYNLGG GAQIITVGRE 180
181 LHDGHWHKVQ VLRNDEQTSL IVDGVSQQRS TKGKEFQFGK FASNSDVYVG GMPNWYSSKL 240
241 ALLALPSVIF EPRFRGAIRN LVYADQPGGS TRRQEIKQQR DIKCGDVPCD HGELPARERP 300
301 LRGVRGGNTT DACERNDPCQ HGGICISTDS GPICECRNLE YDGQYCEKEK APSEATFRGT 360
361 QFLSYDLGQT GAEPIVSAQD AISFYFRTRQ PNGLLFYTGH GTDYLNLALR DGGVSLTMGL 420
421 ANGKQEMHIK PSKVRFDDHQ WHKVTVHRRI QEISSITSFC RLVTVVDDVY TDHSHIAGKF 480
481 TMLSSSRVYV GGAVNPRALL GARVHTNFVG CLRKVEFSAD TLNLNLIDLA KSGSKLIQVA 540
541 GNLEYQCPSG DPQDPVTFTT RESHLVLPPW ETGKQSSISF KFRTKEPNGI IVLATGSKQP 600
601 RAKNPVLIAI ELLNGHIYIH LDLGSGASKV RASRRRVDDG DWHDLILRRN GRDAKVSVDG 660
661 VWNDFRTPGD GTILELDGHM YLGGVGPAYN SVSWPAAIWT ATLRQGFVGC LRDLVLSGKA 720
721 IDIAAFARVQ DSASVKPSCH VQANVCNGNP CLNGGTCLEG WNRPICDCSA TLYGGPTCGR 780
781 ELATLAFNGS QHMTIWLGNG QGTKTQTEEL VIRFKTSRPA GLLLLTSAES NSPDRLEIAL 840
841 VAGRVRASVR LSDREKNLLA GQSVLNDNNW HTIRFSRRAS NLRLQVDGAP PVRAETILGR 900
901 HSTMEIRSVH LGGLFHAEEE IQMTSTMPNF VGQMQGLVFN GQRYLDIVKS LGPELSALPS 960
961 ATFKLTARFV NSPAGQPYHA ATFRSKHSYV GLAMLKAYNS ISIDFRFKTV EPNGLLVFNG1020
1021 GRRNDFVAVE LVNGHIHYTF DLGDGPVTMR DKSRIHMNDN RWHQVSIRRP GPKTHTLTVD1080
1081 DSFEIISLTG NNMHLELAGI LYIGGVFKDM YSKLPASISS RSGFEGCLAS LDLGDASPSL1140
1141 TSDAVVPSSL VVSGCEGPTK CSQNACANRG NCVQQWNAYA CECDMTSYTG PTCYDESIAY1200
1201 EFGNNKGMVQ YTFPENAQAD TEEDNIALGF ITTRPDAVLL RVESATTQDY MELEIVEGNI1260
1261 FMVYNIGSVD LPLGEIGTKV NDNAYHVVRF QRKGGNATLQ LDDYNVQALT PQSHHSTVFN1320
1321 TMSNVQVGGK FSRNGRNRIE RPFAGVIAGL SVNKLRILDL AVERDPHITI RGDVQLVTGV1380
1381 LDRNDLQRMQ QTPASGYPGA LDDLIFSGAG SGCRGDDEDE CTPPFESGSG DDLITPVYVP1440
1441 PTKQTTTSQQ GNSLSTGGSS SGGVITNGTE RACDDEDCVH GSGDYGETTE QFTSTSTARG1500
1501 SESNNEMVTI TTTGRSDVTT EQHQGSSSSS SSGSTPSYAT TQSSSSSSSG GSAASTPGQV1560
1561 STTSSVATSS TQRATSSSTS TSSSTTSTTT TTTTTQATPP PEIRSTVTER ETTPYDIYIA1620
1621 GGGMGGSGRN DHDRMQLPDE HHPLPPLPPP IPPQDPPPYG PYGGNTYNTN MNNYRPKGKG1680
1681 GRINSIEEER TAMIIGIVAG ILIAVVLVIL LVLWLKSNGD RGYKTESEKA AAYGSHNPNA1740
1741 ALLGNTSTNG SYHQQRQHHM HGGGGGGGAG QQQHHAQQQM HNGHNGNGNG GGGGGGGMMS1800
1801 SGSGSLGYGS DGRPQMAGLV QPKAKKRDSK DVKEWYV |
SHOWING SINGLE HITS. [ Hide Single Hits ]
New Feature: Upload Your Own Microscopy Data
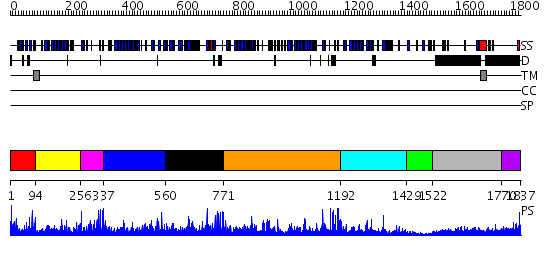
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..93] | N/A | No confident structure predictions are available. | |
| 2 | View Details | [94..255] | 8.69897 | Sex hormone-binding globulin | |
| 3 | View Details | [256..336] | 2.2 | NMR structure of the human NOTCH-1 ligand binding region | |
| 4 | View Details | [337..559] | 31.69897 | No description for 2h0bA was found. | |
| 5 | View Details | [560..770] | 4.69897 | Low density lipoprotein (LDL) receptor YWTD domain; Low density lipoprotein (LDL) receptor, different EGF domains; Ligand-binding domain of low-density lipoprotein receptor | |
| 6 | View Details | [771..1191] | 51.39794 | No description for 2jd4A was found. | |
| 7 | View Details | [1192..1428] | 24.69897 | Ligand-binding domain of neurexin 1beta | |
| 8 | View Details | [1429..1521] | N/A | No confident structure predictions are available. | |
| 9 | View Details | [1522..1769] | 2.144996 | View MSA. No confident structure predictions are available. | |
| 10 | View Details | [1770..1837] | 1.09 | Structure of S5a bound to monoubiquitin provides a model for polyubiquitin recognition |
Functions predicted (by domain):
| # | Gene Ontology predictions | |||||||||||||||||||||||||||||||||||||||||||||
| 1 | No functions predicted. | |||||||||||||||||||||||||||||||||||||||||||||
| 2 | No functions predicted. | |||||||||||||||||||||||||||||||||||||||||||||
| 3 | No functions predicted. | |||||||||||||||||||||||||||||||||||||||||||||
| 4 | No functions predicted. | |||||||||||||||||||||||||||||||||||||||||||||
| 5 | No functions predicted. | |||||||||||||||||||||||||||||||||||||||||||||
| 6 | No functions predicted. | |||||||||||||||||||||||||||||||||||||||||||||
| 7 |
|
|||||||||||||||||||||||||||||||||||||||||||||
| 8 | No functions predicted. | |||||||||||||||||||||||||||||||||||||||||||||
| 9 | No functions predicted. | |||||||||||||||||||||||||||||||||||||||||||||
| 10 | No functions predicted. |
| Source: Reynolds et al. 2008. Manuscript submitted |

|