













| General Information: |
|
| Name(s) found: |
KHDRBS1
[HGNC (HUGO)]
|
| Description(s) found:
Found 37 descriptions. SHOW ALL |
|
| Organism: | Homo sapiens |
| Length: | 443 amino acids |
Gene Ontology: |
|
| Cellular Component: | NONE FOUND |
| Biological Process: | NONE FOUND |
| Molecular Function: | NONE FOUND |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MQRRDDPAAR MSRSSGRSGS MDPSGAHPSV RQTPSRQPPL PHRSRGGGGG SRGGARASPA 60
61 TQPPPLLPPS ATGPDATVGG PAPTPLLPPS ATASVKMEPE NKYLPELMAE KDSLDPSFTH 120
121 AMQLLTAEIE KIQKGDSKKD DEENYLDLFS HKNMKLKERV LIPVKQYPKF NFVGKILGPQ 180
181 GNTIKRLQEE TGAKISVLGK GSMRDKAKEE ELRKGGDPKY AHLNMDLHVF IEVFGPPCEA 240
241 YALMAHAMEE VKKFLVPDMM DDICQEQFLE LSYLNGVPEP SRGRGVPVRG RGAAPPPPPV 300
301 PRGRGVGPPR GALVRGTPVR GAITRGATVT RGVPPPPTVR GAPAPRARTA GIQRIPLPPP 360
361 PAPETYEEYG YDDTYAEQSY EGYEGYYSQS QGDSEYYDYG HGEVQDSYEA YGQDDWNGTR 420
421 PSLKAPPARP VKGAYREHPY GRY |
The following runs contain data for this protein:
| BAIT | COMMENTS | PUBLICATION | |
| View Run | No Comments | Cheeseman, I.M., et al. (2004) | |
| View Run | No Comments | Cheeseman, I.M., et al. (2004) |
SHOWING SINGLE HITS. [ Hide Single Hits ]
New Feature: Upload Your Own Microscopy Data
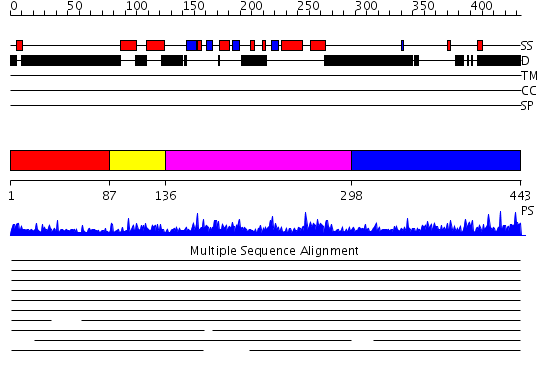
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..86] | N/A | No confident structure predictions are available. | |
| 2 | View Details | [87..135] | 3.118997 | View MSA. No confident structure predictions are available. | |
| 3 | View Details | [136..297] | 53.69897 | Solution structure of the KH-QUA2 region of the Xenopus STAR-GSG Quaking protein. | |
| 4 | View Details | [298..443] | 3.104997 | View MSA. No confident structure predictions are available. |
Functions predicted (by domain):
| # | Gene Ontology predictions | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 1 | No functions predicted. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 2 | No functions predicted. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 3 |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 4 | No functions predicted. |
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.99 |
Source: Reynolds et al. (2008)