













| General Information: |
|
| Name(s) found: |
gi|24375059
[NCBI NR]
gi|24349807 [NCBI NR] |
| Description(s) found:
Found 5 descriptions. SHOW ALL |
|
| Organism: | Shewanella oneidensis MR-1 |
| Length: | 365 amino acids |
Gene Ontology: |
|
| Cellular Component: | NONE FOUND |
| Biological Process: |
purine ribonucleotide biosynthetic process
[ISS |
| Molecular Function: |
phosphoribosylaminoimidazole carboxylase activity
[ISS |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MTQTSTKPKV WVLGNGQLGA MLTHAGEPLA IDVRAVDIMT PTDDILPLAP NDIITAEREQ 60
61 WPESALSLQL STHPHFVNGP VFSRLADRYS QKSLLDQLNV PTAPWSLVDD HTKVETLYQA 120
121 FGPRVLMKRR TGGYDGKGQH WLKQAEAGDI PHDWRNLAIA EQAINFDEEV SLVGVRTREG 180
181 QCVFYPLTLN LHQDGILMAS IAPLARLDHL QAQAETMLSA IMHELEYAGV MAMECFRVGD 240
241 NLLVNELAPR VHNSGHWTQA GTHMDQFQLH LRALCGIAIP QPQVNFQCVM VNLIGIDNDP 300
301 RWLSLPNAEL YWYNKEVRPG RKVGHLNLSV PNLTVLTNSI SALQTWMPNQ YQAPLAWILA 360
361 EFTKS |
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
New Feature: Upload Your Own Microscopy Data
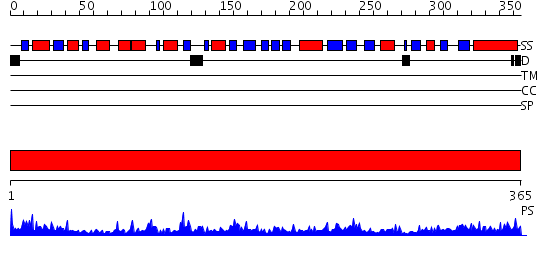
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..365] | 62.522879 | N5-carboxyaminoimidazole ribonucleotide synthetase PurK (AIRC), C-domain; N5-carboxyaminoimidazole ribonucleotide synthetase PurK (AIRC), N-domain; N5-carboxyaminoimidazole ribonucleotide synthetase PurK (AIRC), domain 2 |
Functions predicted (by domain):
| # | Gene Ontology predictions | ||||||||||||||||||||||||||||||||||||
| 1 |
|
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.73 |
Source: Reynolds et al. (2008)