













| General Information: |
|
| Name(s) found: |
gi|7494332
[NCBI NR]
|
| Description(s) found:
Found 2 descriptions. SHOW ALL |
|
| Organism: | Plasmodium falciparum |
| Length: | 539 amino acids |
Gene Ontology: |
|
| Cellular Component: |
chaperonin-containing T-complex
[ISS |
| Biological Process: | NONE FOUND |
| Molecular Function: | NONE FOUND |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MSHLMSLPIV LLKEGTDTAQ GRSQIIRNIN ACQIIVDIVK TTLGPRGMDK LIYTERDVTI 60
61 TNDGATVMNL LNISHPAASI LVDIAKSQDD EVGDGTTSVV VVAGELLNEA KGLLNDGIEP 120
121 NMIIDGFRNA CNVAINKLNE LSLNFSNKNE EEKRSILLKC AQTALNSKLV SNHKEFFGEL 180
181 VVNAVYKLGD NLDKSNIGIK KVTGGSCLDT QLIYGVAFKK TFSYAGFEQQ PKKFINPKIL 240
241 LLNVELELKA EKENAEVRIE NPNEYNSIVQ AEWDIIFKKL NLIKDCGANI VLSKLPIGDI 300
301 ATQFFADHDI FCAGRVEDAD LKRTANATGA LVQTSLFNLN DDVLGTCGVF EEVQIGNERY 360
361 NIFKECLKTK SVTIILRGGA KQFIEEVERS INDAIMIVLR CITNSEIVPG AGSIEMQLSK 420
421 YLRIYSRSIC NKEQIVLFSF AKALESIPRH LSHNAGYDST DILNKLRKKH SEQTSDIWYG 480
481 VDCMEGDIIN AYDNCIFEVT KIKRNVIYSA TEAACLILSI DETIKNPSSA AGTQRSPYS |
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
New Feature: Upload Your Own Microscopy Data
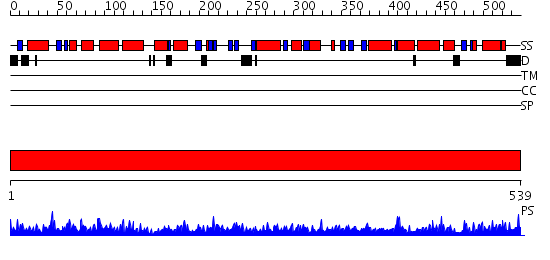
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..539] | 151.0 | Crystal structure of the chaperonin from Thermococcus strain KS-1 (nucleotide-free form of single mutant) |
Functions predicted (by domain):
| # | Gene Ontology predictions | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 1 |
|
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.59 |
Source: Reynolds et al. (2008)