













| General Information: |
|
| Name(s) found: |
gi|4567234
[NCBI NR]
gi|25286034 [NCBI NR] gi|15226577 [NCBI NR] |
| Description(s) found:
Found 11 descriptions. SHOW ALL |
|
| Organism: | Arabidopsis thaliana |
| Length: | 492 amino acids |
Gene Ontology: |
|
| Cellular Component: | NONE FOUND |
| Biological Process: |
nucleotide biosynthetic process
[IEA]
dTMP biosynthetic process [IEA] glycine biosynthetic process [IEA] one-carbon metabolic process [IEA] |
| Molecular Function: |
thymidylate synthase activity
[IEA]
dihydrofolate reductase activity [IEA] |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MASTGETICN NGQSAVSVTR RRSYQVVIAA TRDMGLGMDM KLPWDLPSEY QFFQDVTTRT 60
61 SDPTKRNATI MGRKSWESTP LEIRPLPGRL NIVLTKSSCH NIAIDENVLV SSSMESALEL 120
121 LATEPYSLSI EKVFVIGGGE LLRNYMNASI CDAIHLTEID ISVPCDAFAP RVDTSLYRPW 180
181 YSSFPVVENG IRYSFNTYVR RKDAIVGSGE KKSVAESDLK EYSFLPKMVF ERHEEFGYLN 240
241 LVQNIISSGD MNDNSTLSKF GCQMRFNLRK TFPLLTTKKI FWLGVVEEIL QLISGSNNPK 300
301 ENGSHIWDTD EAKEYLDSFG VNATEEDGDN PFLHGLHWKH CDARFVIQEF SQLSDVINKI 360
361 KNNPHDQRIM LAACNPLDFK LSVSPCHTFT QFYVANGEVS CQIYQSSTEA SIGIPFSIAT 420
421 YSLLTCIIAH VCDLGAGDFI HVIGQAYINK AHVKAIQKQL QISPKPFPIL KINPEKKKMD 480
481 NFEASDLELM RI |
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
New Feature: Upload Your Own Microscopy Data
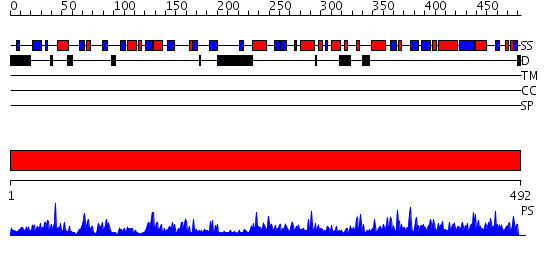
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..492] | 152.0 | Crystal structure of DHFR-TS from Cryptosporidium hominis |
Functions predicted (by domain):
| # | Gene Ontology predictions | |||||||||||||||||||||||||||||||||||||||
| 1 |
|
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.85 |
Source: Reynolds et al. (2008)