













| General Information: |
|
| Name(s) found: |
SPPA1_ARATH
[Swiss-Prot]
|
| Description(s) found:
Found 11 descriptions. SHOW ALL |
|
| Organism: | Arabidopsis thaliana |
| Length: | 677 amino acids |
Gene Ontology: |
|
| Cellular Component: |
chloroplast
[IDA]
chloroplast photosystem II [IEP] chloroplast thylakoid membrane [IDA][IEP] |
| Biological Process: |
proteolysis
[IDA]
response to light intensity [IEP] |
| Molecular Function: |
serine-type endopeptidase activity
[IDA]
|
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MAKLLLLHAP HVIPRFSSSS SRSLVSAAAL YRRPLLVNPQ FSHIGPRLHS PYNRRFSARA 60
61 FDDSPASSAE MEKEKQEQLL DGVSGKKDED YPTGEMEYEN RNAWEIFVVK FRMLFAYPWQ 120
121 RVRKGSVLTM TLRGQISDQL KSRFNSGLSL PQLSENFVKA AYDPRIAGVY LHIDPLSCGW 180
181 GKVEEIRRHI LNFKKSGKFI VGYISICGLK EYYLGCACNE LFAPPSAYSF LYGLTVQASF 240
241 LGGVFEKVGI EPQVQRIGKY KSAGDQLSRK SISEENYEML SVLLDNIYSN WLDGVSDATG 300
301 KKREDVENFI NQGVYEIEKL KEAGLIKDIR YDDEVITMLK ERLGVEKDKK LPTVDYKKYS 360
361 GVKKWTLGLT GGRDQIAIIR AGGSISRVKG PLSTPGSAII AEQLIEKIRS VRESKKYKAA 420
421 IIRIDSPGGD ALASDLMWRE IKLLAETKPV IASMSDVAAS GGYYMAMAAN AIVAENLTLT 480
481 GSIGVVTARF TLAKLYEKIG FNKETISRGK YAELLGAEER PLKPEEAELF EKSAQHAYQL 540
541 FRDKAALSRS MPVDKMEEVA QGRVWTGKDA HSRGLIDAVG GLSRAIAIAK QKANIPLNKK 600
601 VTLVELSRPS TSLPDILSGI GSSVIGVDRT LKGLLDELTI TEGVQARMDG IMFQQLGRDS 660
661 LATPIIDMLK DYLSSLR |
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
New Feature: Upload Your Own Microscopy Data
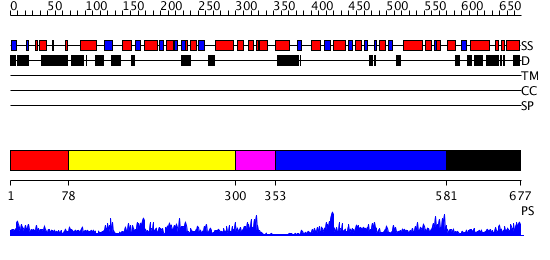
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..77] | N/A | No confident structure predictions are available. | |
| 2 | View Details | [78..299] | 3.221849 | Crystal Structure of Enoyl-CoA Hydratase from Thermus Thermophilus HB8 | |
| 3 | View Details | [300..352] | 2.12 | Crystal structure of the ATP-dependent Clp Protease proteolytic subunit 1 (ClpP1) from Mycobacterium tuberculosis | |
| 4 | View Details | [353..580] | 44.69897 | Clp protease, ClpP subunit | |
| 5 | View Details | [581..677] | N/A | Confident ab initio structure predictions are available. |
| Source: Reynolds et al. 2008. Manuscript submitted |

|