













| General Information: |
|
| Name(s) found: |
frs-1 /
CE33173
[WormBase]
|
| Description(s) found:
Found 16 descriptions. SHOW ALL |
|
| Organism: | Caenorhabditis elegans |
| Length: | 496 amino acids |
Gene Ontology: |
|
| Cellular Component: |
cytoplasm
[IEA |
| Biological Process: |
tRNA aminoacylation for protein translation
[IEA nematode larval development [IMP positive regulation of growth rate [IMP embryonic development ending in birth or egg hatching [IMP post-embryonic body morphogenesis [IMP phenylalanyl-tRNA aminoacylation [IEA reproduction [IMP growth [IMP translation [IEA |
| Molecular Function: |
nucleotide binding
[IEA ATP binding [IEA aminoacyl-tRNA ligase activity [IEA phenylalanine-tRNA ligase activity [IEA |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MTAETDVRST ENLPQQILDF LQESNEFNSI QLAQQWNLDH QKVIGAIKSL LANEGVLTTK 60
61 DVTEKRLELT NEGVQFANEG SPEYLVFEFV GTDGAAQADI QKKPFGKIGM AKAMQFKWVS 120
121 VDKGRVVRQA TEVTDSTRKQ LESLRIGSSD VSENEKKELK KRKLISEVNI KALVVSKGTS 180
181 FTTSLAKQEA DLTPEMIASG SWKDMQFKKY NFDSLGVVPS SGHLHPLMKV RSEFRQIFFS 240
241 MGFSEMATNR YVESSFWNFD ALFQPQQHPA RDAHDTFFVS DPAISTKFPE DYLERVKTVH 300
301 SKGGYGSAGY NYDWKIEEAQ KNVLRTHTTA VSARQLYQLA QEGFRPSKLF SIDRVFRNET 360
361 LDATHLAEFH QVEGVIAEKN LSLAHLIGIF TEFFKKLGIT NLRFKPTYNP YTEPSMEIFA 420
421 YHQGLTKWVE IGNSGMFRPE MLLPMGLPAD VNVAGYGLSL ERPTMIKYGI NNIRDLFGSK 480
481 IDLNVVYNNP ICRLDK |
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
New Feature: Upload Your Own Microscopy Data
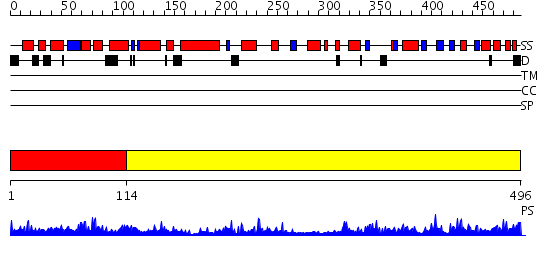
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..113] | 1.15 | No description for 2gqqA was found. | |
| 2 | View Details | [114..496] | 74.09691 | Phenyl-tRNA synthetase (PheRS) alpha subunit, PheS |
Functions predicted (by domain):
| # | Gene Ontology predictions | ||||||||||||||||||||||||||||||||||||||||||||||||
| 1 | No functions predicted. | ||||||||||||||||||||||||||||||||||||||||||||||||
| 2 |
|
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.98 |
Source: Reynolds et al. (2008)