













| General Information: |
|
| Name(s) found: |
cct-2 /
CE16437
[WormBase]
|
| Description(s) found:
Found 22 descriptions. SHOW ALL |
|
| Organism: | Caenorhabditis elegans |
| Length: | 529 amino acids |
Gene Ontology: |
|
| Cellular Component: | NONE FOUND |
| Biological Process: |
hermaphrodite genitalia development
[IMP]
nematode larval development [IMP] embryonic development ending in birth or egg hatching [IMP] cellular protein metabolic process [IEA protein folding [IEA receptor-mediated endocytosis [IMP] reproduction [IMP] growth [IMP] |
| Molecular Function: |
ATP binding
[IEA protein binding [IEA unfolded protein binding [IEA |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MLPVQILKDN AQEERGESAR LSSFVGAIAI GDLVKSTLGP KGMDKILISG NPESAGGIKV 60
61 TNDGATILKS IGVDNPAAKV LVDMSMTQDH EVGDGTTSVT VLAAELLKEA EKLVNQRIHP 120
121 QTIISGYRRA LGIAQESLKK SSIESGDNIR DDLLKIARTT LGSKILSQHK EHFAQLAVDA 180
181 VLRLKGSGNL DAIQIIKKLG GSMNESYLDE GFLLEKLPGM FQPRRVEKAK ILIANTPMDT 240
241 DKVKVFGSRV RVDGVAKVAE LEAAEKLKMK EKVDKILAHN CNVFINRQLI YNYPEQLFAD 300
301 AKVMAIEHAD FEGIERLALV LGGEIVSTFD SPQTAQFGSC DLIEEIMIGE DRLLRFSGVK 360
361 LGEACSVVLR GATQQILDES ERSLHDALCV LVTHVKESKT VAGAGASEIL MSSAIAVEAQ 420
421 KVAGKEALAV EAFGRALAQL PTIICDNAGL DSAELVTRLR AEHANGRHNM GIDIEKGEVA 480
481 DVTKLGVIES YNVKLCMVSS AAEATEQILR VDDIIKAAPR ARAQDNRPC |
The following runs contain data for this protein:
| BAIT | COMMENTS | PUBLICATION | |
| View Run | No Comments | Cheeseman, I.M., et al. (2004) |
SHOWING SINGLE HITS. [ Hide Single Hits ]
New Feature: Upload Your Own Microscopy Data
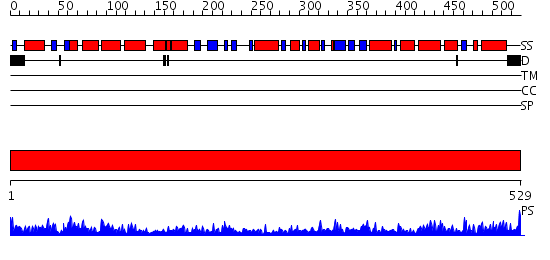
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..529] | 142.0 | Crystal structure of the chaperonin from Thermococcus strain KS-1 (nucleotide-free form of single mutant) |
Functions predicted (by domain):
| # | Gene Ontology predictions | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 1 |
|
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.98 |
Source: Reynolds et al. (2008)