













| General Information: |
|
| Name(s) found: |
bam-PA /
FBpp0084243
[FlyBase]
|
| Description(s) found:
Found 32 descriptions. SHOW ALL |
|
| Organism: | Drosophila melanogaster |
| Length: | 442 amino acids |
Gene Ontology: |
|
| Cellular Component: |
cytoplasm
[IDA][TAS]
fusome [IDA][TAS] spectrosome [IDA][TAS] |
| Biological Process: |
germ cell development
[TAS]
cystoblast division [IMP][TAS] spermatogenesis [TAS] male meiosis [TAS] spermatogonial cell division [TAS] fusome organization [IMP] germ-line stem cell division [IMP][TAS] vesicle targeting to fusome [TAS] gamete generation [IMP][NAS] female germ-line stem cell division [IMP] germarium-derived female germ-line cyst formation [IMP] oogenesis [IMP] female germ-line cyst formation [TAS] cell fate specification [TAS] cell fate determination [TAS] spectrosome organization [TAS] |
| Molecular Function: |
translation repressor activity, nucleic acid binding
[IDA]
|
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MLNARDVCPE GNDDQQLDHN FKQMEEHLAL MVEGNENEDP RKATCEYEDT NEDGATCTSG 60
61 VLSEIQENFG RLRLCDVTAP LLEFHGLDCL QQIQKRSRHF AFDGSPAKKS RSGGVLVTGP 120
121 KQKQLQKENV WNRKSKGSAS ADNIEKLPIT IEKLHMIGLH GDCLEHNAVL RLMNLFRSLH 180
181 DHLTADLGFS RQNSMPSDYL FDMPVKSTMP KSLNVRYQLQ VLCTKVERFL VQQRRTLEAN 240
241 RHFDFEKYDE CDKLLKGFAS YLDNFKLLLK PKMRNRNGNS GSNADKFHTQ RMERLLIGLR 300
301 DWIKAAHLSV HVFNWEMDLE HRYSGAMTES HKSLNERAIL LSGAELRAAE ARGISAEDLF 360
361 IAQRYKLGGP IYCVLEQHEF LSALIANPET YFPPSVVAIC GPQKLGAVSM EQPSASEEEF 420
421 EETEEVPSSP PRHTGRVPRF RS |
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
New Feature: Upload Your Own Microscopy Data
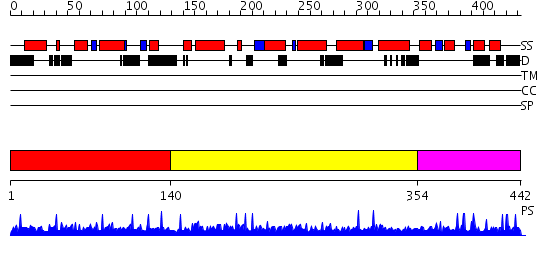
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..139] | N/A | No confident structure predictions are available. | |
| 2 | View Details | [140..353] | N/A | No confident structure predictions are available. | |
| 3 | View Details | [354..442] | N/A | No confident structure predictions are available. |
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.99 |
Source: Reynolds et al. (2008)