













| General Information: |
|
| Name(s) found: |
gi|16331500
[NCBI NR]
gi|1001156 [NCBI NR] |
| Description(s) found:
Found 5 descriptions. SHOW ALL |
|
| Organism: | Synechocystis sp. PCC 6803 |
| Length: | 448 amino acids |
Gene Ontology: |
|
| Cellular Component: | NONE FOUND |
| Biological Process: | NONE FOUND |
| Molecular Function: | NONE FOUND |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MQFAKILIAN RGEIALRIIH SCEELGIPTV AVHSTIDRHA LHVQLANESV CIGPPPSNKS 60
61 YLNIPNIIAA ALTRNATAIH PGYGFLAENA RFAEICADHQ ITFIGPSPEA ITAMGDKSTA 120
121 KKTMQKSGVP CVPGSPGLIE SEATALKIAA EIGYPVIIKA TAGGGGRGMR LVQEEKDFLK 180
181 LFHAAQGEAG AAFGNPGVYL EKFIEKPRHI EFQILADSHG NVVHLGERDC SIQRRHQKLL 240
241 EEAPSPFLTP HLRKKMGEAA VKAAKSINYV GAGTVEFLVD GNGNFYFMEM NTRIQVEHPV 300
301 TEMITGYDLI SEQIRIAMGE KLRFRQSDVE IRGHAIECRI NAEDPKQNFR PHPGKISAYL 360
361 PPGGPGVRID SHVYTDYEIP PYYDSLIGKL IVWAGDRPSA IKRMQRALRE CAITGVPTTL 420
421 EFHQRILQTP AFLAGDVYTN FIEEHLTP |
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
New Feature: Upload Your Own Microscopy Data
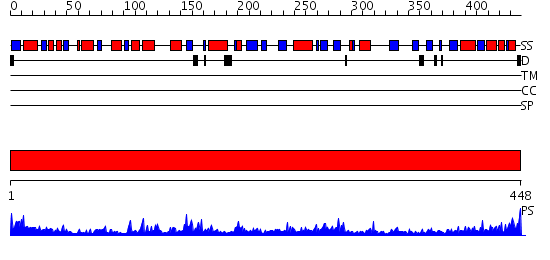
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..448] | 126.0 | Crystal structure of the biotin carboxylase subunit of pyruvate carboxylase |
Functions predicted (by domain):
| # | Gene Ontology predictions | ||||||||||||||||||||||||||||||||||||||||||
| 1 |
|
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.98 |
Source: Reynolds et al. (2008)