













| General Information: |
|
| Name(s) found: |
UGE5_ARATH
[Swiss-Prot]
|
| Description(s) found:
Found 17 descriptions. SHOW ALL |
|
| Organism: | Arabidopsis thaliana |
| Length: | 351 amino acids |
Gene Ontology: |
|
| Cellular Component: |
endomembrane system
[IEA]
|
| Biological Process: |
response to stress
[IMP]
|
| Molecular Function: |
UDP-glucose 4-epimerase activity
[IDA]
protein dimerization activity [IPI] |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MMARNVLVSG GAGYIGSHTV LQLLLGGYSV VVVDNLDNSS AVSLQRVKKL AAEHGERLSF 60
61 HQVDLRDRSA LEKIFSETKF DAVIHFAGLK AVGESVEKPL LYYNNNLVGT ITLLEVMAQH 120
121 GCKNLVFSSS ATVYGSPKEV PCTEEFPISA LNPYGRTKLF IEEICRDVYG SDPEWKIILL 180
181 RYFNPVGAHP SGDIGEDPRG IPNNLMPFVQ QVAVGRRPHL TVFGNDYNTK DGTGVRDYIH 240
241 VIDLADGHIA ALRKLEDCKI GCEVYNLGTG NGTSVLEMVD AFEKASGKKI PLVIAGRRPG 300
301 DAEVVYASTE RAESELNWKA KYGIEEMCRD LWNWASNNPY GYDSSSEDNS H |
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
New Feature: Upload Your Own Microscopy Data
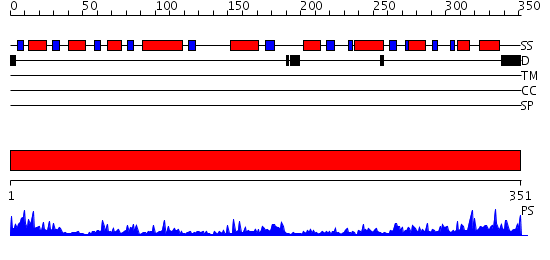
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..351] | 90.045757 | Uridine diphosphogalactose-4-epimerase (UDP-galactose 4-epimerase) |
Functions predicted (by domain):
| # | Gene Ontology predictions | |||||||||||||||||||||||||||||||||||||||||||||
| 1 |
|
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.55 |
Source: Reynolds et al. (2008)