













| General Information: |
|
| Name(s) found: |
KUP_ECOLI
[Swiss-Prot]
|
| Description(s) found:
Found 8 descriptions. SHOW ALL |
|
| Organism: | Escherichia coli |
| Length: | 622 amino acids |
Gene Ontology: |
|
| Cellular Component: | NONE FOUND |
| Biological Process: | NONE FOUND |
| Molecular Function: | NONE FOUND |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MSTDNKQSLP AITLAAIGVV YGDIGTSPLY TLRECLSGQF GFGVERDAVF GFLSLIFWLL 60
61 IFVVSIKYLT FVMRADNAGE GGILTLMSLA GRNTSARTTS MLVIMGLIGG SFFYGEVVIT 120
121 PAISVMSAIE GLEIVAPQLD TWIVPLSIIV LTLLFMIQKH GTAMVGKLFA PIMLTWFLIL 180
181 AGLGLRSIIA NPEVLHALNP MWAVHFFLEY KTVSFIALGA VVLSITGVEA LYADMGHFGK 240
241 FPIRLAWFTV VLPSLTLNYF GQGALLLKNP EAIKNPFFLL APDWALIPLL IIAALATVIA 300
301 SQAVISGVFS LTRQAVRLGY LSPMRIIHTS EMESGQIYIP FVNWMLYVAV VIVIVSFEHS 360
361 SNLAAAYGIA VTGTMVLTSI LSTTVARQNW HWNKYFVALI LIAFLCVDIP LFTANLDKLL 420
421 SGGWLPLSLG TVMFIVMTTW KSERFRLLRR MHEHGNSLEA MIASLEKSPP VRVPGTAVYM 480
481 SRAINVIPFA LMHNLKHNKV LHERVILLTL RTEDAPYVHN VRRVQIEQLS PTFWRVVASY 540
541 GWRETPNVEE VFHRCGLEGL SCRMMETSFF MSHESLILGK RPWYLRLRGK LYLLLQRNAL 600
601 RAPDQFEIPP NRVIELGTQV EI |
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
New Feature: Upload Your Own Microscopy Data
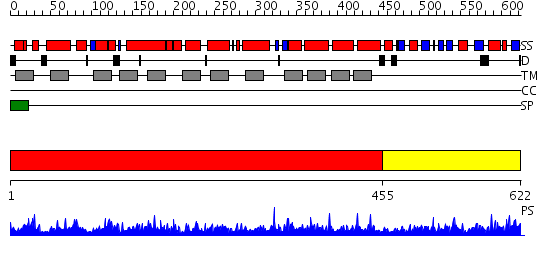
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..454] | 1.21 | Crystal structure of LEUTAA, a bacterial homolog of Na+/Cl--dependent neurotransmitter transporters | |
| 2 | View Details | [455..622] | 3.853872 | No description for PF02705.7 was found. No confident structure predictions are available. |
Functions predicted (by domain):
| # | Gene Ontology predictions | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 1 |
|
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 2 | No functions predicted. |
| Source: Reynolds et al. 2008. Manuscript submitted |

|