













| General Information: |
|
| Name(s) found: |
EFGM1_ARATH
[Swiss-Prot]
|
| Description(s) found:
SHOW ONLY BEST |
|
| Organism: | Arabidopsis thaliana |
| Length: | 754 amino acids |
Gene Ontology: |
|
| Cellular Component: |
mitochondrion
[IDA]
|
| Biological Process: |
response to cadmium ion
[IEP]
translational elongation [IEA] |
| Molecular Function: |
GTP binding
[IEA]
ATP binding [IDA] translation elongation factor activity [IEA][ISS] GTPase activity [IEA] translation factor activity, nucleic acid binding [ISS] |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MARFPTSPAP NRLLRLFSSN KRSSSPTAAL LTGDFQLIRH FSAGTAARVA KDEKEPWWKE 60
61 SMDKLRNIGI SAHIDSGKTT LTERVLFYTG RIHEIHEVRG RDGVGAKMDS MDLEREKGIT 120
121 IQSAATYCTW KDYKVNIIDT PGHVDFTIEV ERALRVLDGA ILVLCSVGGV QSQSITVDRQ 180
181 MRRYEVPRVA FINKLDRMGA DPWKVLNQAR AKLRHHSAAV QVPIGLEENF QGLIDLIHVK 240
241 AYFFHGSSGE NVVAGDIPAD MEGLVAEKRR ELIETVSEVD DVLAEKFLND EPVSASELEE 300
301 AIRRATIAQT FVPVFMGSAF KNKGVQPLLD GVVSFLPSPN EVNNYALDQN NNEERVTLTG 360
361 SPDGPLVALA FKLEEGRFGQ LTYLRVYEGV IKKGDFIINV NTGKRIKVPR LVRMHSNDME 420
421 DIQEAHAGQI VAVFGIECAS GDTFTDGSVK YTMTSMNVPE PVMSLAVQPV SKDSGGQFSK 480
481 ALNRFQKEDP TFRVGLDPES GQTIISGMGE LHLDIYVERM RREYKVDATV GKPRVNFRET 540
541 ITQRAEFDYL HKKQSGGAGQ YGRVTGYVEP LPPGSKEKFE FENMIVGQAI PSGFIPAIEK 600
601 GFKEAANSGS LIGHPVENLR IVLTDGASHA VDSSELAFKM AAIYAFRLCY TAARPVILEP 660
661 VMLVELKVPT EFQGTVAGDI NKRKGIIVGN DQEGDDSVIT ANVPLNNMFG YSTSLRSMTQ 720
721 GKGEFTMEYK EHSAVSNEVQ AQLVNAYSAS KATE |
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
New Feature: Upload Your Own Microscopy Data
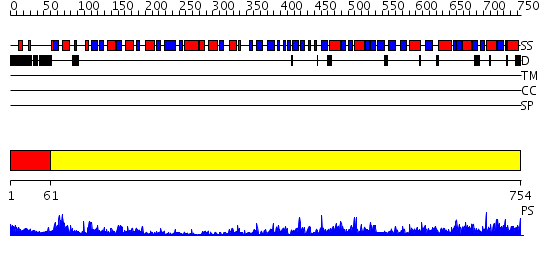
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..60] | 7.522879 | Probable GTPase Der, N-terminal and middle domains; Probable GTPase Der, C-terminal domain | |
| 2 | View Details | [61..754] | 1000.0 | Elongation factor G (EF-G), domain II; Elongation factor G (EF-G), N-terminal (G) domain; Elongation factor G (EF-G), domain IV; Elongation factor G (EF-G) |
Functions predicted (by domain):
| # | Gene Ontology predictions | ||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 1 | No functions predicted. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 2 |
|
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.54 |
Source: Reynolds et al. (2008)