













| General Information: |
|
| Name(s) found: |
ULA1_ARATH
[Swiss-Prot]
|
| Description(s) found:
Found 18 descriptions. SHOW ALL |
|
| Organism: | Arabidopsis thaliana |
| Length: | 540 amino acids |
Gene Ontology: |
|
| Cellular Component: |
nucleus
[IEP]
|
| Biological Process: |
DNA repair
[IMP]
auxin homeostasis [NAS] leaf morphogenesis [IGI] auxin mediated signaling pathway [IMP] protein ubiquitination [IGI] response to water deprivation [IMP] |
| Molecular Function: |
small protein activating enzyme activity
[IDA][ISS]
|
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MQAVKRSRRH VEEEPTMVEP KTKYDRQLRI WGEVGQAALE EASICLLNCG PTGSEALKNL 60
61 VLGGVGSITV VDGSKVQFGD LGNNFMVDAK SVGQSKAKSV CAFLQELNDS VNAKFIEENP 120
121 DTLITTNPSF FSQFTLVIAT QLVEDSMLKL DRICRDANVK LVLVRSYGLA GFVRISVKEH 180
181 PIIDSKPDHF LDDLRLNNPW PELKSFVETI DLNVSEPAAA HKHIPYVVIL VKMAEEWAQS 240
241 HSGNLPSTRE EKKEFKDLVK SKMVSTDEDN YKEAIEAAFK VFAPRGISSE VQKLINDSCA 300
301 EVNSNSSAFW VMVAALKEFV LNEGGGEAPL EGSIPDMTSS TEHYINLQKI YLAKAEADFL 360
361 VIEERVKNIL KKIGRDPSSI PKPTIKSFCK NARKLKLCRY RMVEDEFRNP SVTEIQKYLA 420
421 DEDYSGAMGF YILLRAADRF AANYNKFPGQ FDGGMDEDIS RLKTTALSLL TDLGCNGSVL 480
481 PDDLIHEMCR FGASEIHVVS AFVGGIASQE VIKLVTKQFV PMLGTYIFNG IDHKSQLLKL 540
541 |
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
New Feature: Upload Your Own Microscopy Data
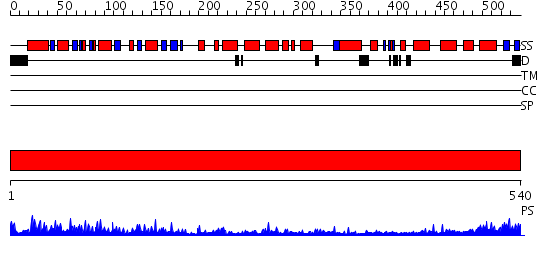
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..540] | 105.0 | Structure of APPBP1-UBA3-Ubc12N26: a unique E1-E2 interaction required for optimal conjugation of the ubiquitin-like protein NEDD8 |
Functions predicted (by domain):
| # | Gene Ontology predictions | |||||||||||||||||||||||||||||||||||||||||||||
| 1 |
|
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.98 |
Source: Reynolds et al. (2008)