













| General Information: |
|
| Name(s) found: |
SPBC1289.10c
[Sanger Pombe]
|
| Description(s) found:
Found 19 descriptions. SHOW ALL |
|
| Organism: | Schizosaccharomyces pombe |
| Length: | 743 amino acids |
Gene Ontology: |
|
| Cellular Component: |
nucleus
[ISS cytoplasm [IDA |
| Biological Process: |
positive regulation of cell-substrate adhesion
[IMP positive regulation of gene-specific transcription from RNA polymerase II promoter [ISS] gene-specific transcription from RNA polymerase II promoter [IC |
| Molecular Function: |
transcription factor activity
[IC]
specific RNA polymerase II transcription factor activity [ISS single-stranded DNA binding [ISS |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MADPGLRSGV GLPSQQGQKH DLQKDQKQPH VNNADRTTQS LLNSYIYDYL IKKDYCEAAR 60
61 AFGREAQVQT LVRSQEETNS LAKRHKRMSP VAVKHEGISN NESSDENMNV NNGNLDSFSS 120
121 SSAPPPPPIL PIDSAGGFLI EWWNVFWDIY NARRGQGSEP AKAYMSHISN LRKKSRLNLQ 180
181 EIQKNSLHTG NTSHPYANAS FPHDPANAMG QQIDSSQFHQ GAGGLNDRNQ HLMRQAMLNN 240
241 QSRETFPPTA AQLQQLKQLH YRQLQSVQQQ QKQHQQKKTP QSGSTPQMQN TTSQPTTHDT 300
301 HPPKQQGPIS DFRSIPSSPK TEGAPSNAQF RPSLPATPNG SVPQSNPLYD TTGLNGGQYP 360
361 VVQNSAQPLL HEINFASNRN PHLKQGGAVP SSTLPQQQKS LDKPKPAQQP STGQFSGNQM 420
421 NQYGFSNSPY SQNMLYNFNG NANPSRLNPA LKNYMEELKL LEQQNKKRLL LVSQEKERKG 480
481 YTSASPDRPL SQTITESSVA KTKSTTPKST DTPTEATTSP VKVSTKNSNT TENLNGINES 540
541 NMPMLQNGLP LRTSGDHPSN YSNLIENSST SDTNNADNGM DVMGNWQLQQ THSSRPTPNA 600
601 SSPLDVRSKQ KPSSANSNAP TPAPTVNTTN PESSTNEATS VGPALEPSQG ANVHKSDSEL 660
661 DNQNQSGKSN PDTSATPSAP TESTTVATKS SDNQLLDVGN STDIDAALLN DFDFDKFLKD 720
721 TSTGDDLWFG LFNLPDNEDS TAA |
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
New Feature: Upload Your Own Microscopy Data
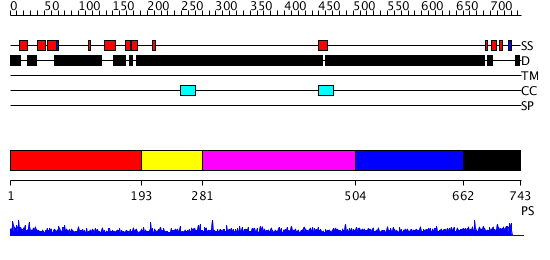
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..192] | N/A | No confident structure predictions are available. | |
| 2 | View Details | [193..280] | N/A | No confident structure predictions are available. | |
| 3 | View Details | [281..503] | N/A | No confident structure predictions are available. | |
| 4 | View Details | [504..661] | N/A | No confident structure predictions are available. | |
| 5 | View Details | [662..743] | N/A | No confident structure predictions are available. |
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.99 |
Source: Reynolds et al. (2008)