













| General Information: |
|
| Name(s) found: |
clr1 /
SPBC2D10.17
[Sanger Pombe]
|
| Description(s) found:
SHOW ONLY BEST |
|
| Organism: | Schizosaccharomyces pombe |
| Length: | 1238 amino acids |
Gene Ontology: |
|
| Cellular Component: |
mating-type region heterochromatin
[IDA nucleus [IDA telomeric heterochromatin [IDA NuRD complex [IDA rDNA heterochromatin [IDA centromeric heterochromatin [IDA |
| Biological Process: |
negative regulation of transcription from RNA polymerase II promoter
[IMP chromatin modification [IEA] chromatin silencing at centromere [IMP chromatin silencing at rDNA [IMP chromatin silencing [IMP chromatin silencing at silent mating-type cassette [IMP |
| Molecular Function: |
zinc ion binding
[IEA]
|
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MAEPDISSSE TLTELQQLRL LYFFCFYHAA PFNVKTLVHS LIPPGALSYL LSTYDILRPW 60
61 LMALRVREGP VNDISTIVQL YEEIVKTGFF INPPPFESYS QTLVARITTL GRPKLQVQQE 120
121 AQSEVYQRAS TNTQQQVSNV SHGNFKPNSS VNTEPNTSIL SNSKYAGIKP FDFQSSSQNP 180
181 GLVCEQKTFY QHQFRPFSNL PSNKSSPVKH VSPNVKNNSK KTASSVNSNH SSIPSSITKS 240
241 NISSLDVYGS EKLISSGSQQ PGHGMVQTTS DKVNASASLY DRSPSKKDIT SSRNTSSYNL 300
301 GSMRNPSTLK NAAHANPFEG LRFQGSSAVL KEGLNSTVKK TFFDNLNSEK VCPSVSPFLT 360
361 PDNIASSILY STASFSRSKP DRPRLNLSLE LKLMQNELNK GQLKKQFKGD LRNLADWNNL 420
421 SLVSSKFPSL PITNLRPDGS FLKHRRFNEE IAYNRQTLEK AIKQLDLSPD KVIQLREQNG 480
481 VAVNGRVCYP TRNKHSEISA QSSSSLGVTK SLASEVYSSS TVDTISKLNT DKDNYLIKSK 540
541 KEPIQQKSVS SETTLVKPSS TSSYIDTTNN VLKTNSSFKS SGLTSGPRNE KELLPEGIPT 600
601 SHNNSETQAQ TADVSNIAAS ADGIYNSDQE KPPEKLDVTK RAFGREIENS NEKELLTSTF 660
661 LSPSAESQVC LAEIKTIRPG LVPKKQFSVD QNNVISDNTD CSLPKPSNSK LSSISSDGDA 720
721 SSNRMAVPDK SPFVHAAPNS KALTKDSFST HISVSSLLHS DNEISPIDST RKDYFTSKDS 780
781 NLQTLKEDAS STKQAKDSGT NDFDKLISGN DVSKNNSGEE QSRSALKPLI SGKLSSCESI 840
841 NLTKDISTVK RKEYFGIEST SSKQPFHDTG SIKIPAKRSF DTIDKDFRSS NIPFADKIKE 900
901 DGGDKNVISS IHITTELPKS MPVEVPTNAG AQSDQSNVVD SESLNLRENI STSVADVSLS 960
961 QAGNEAVLSK KACKPLVLID PFEEKVLKAF NMLSKGYAEY RCQWEGCLAN LHSLENFIKH1020
1021 VLLLHHPKSC SVVKCLWASC DMVLPSEEFE MHLRGHLNNI RLNCEVSNCK KCFSNYEDMF1080
1081 KHLQHSHLPF KFTPESFIKI RNGNVKEEAR RTRNAYTQKS GEVECFMETC TPIAKPAPAN1140
1141 WYPVPPPGFN SSLLSRLTQS NQSKDKIIAA LAKRNVYKSF AGLYDSKGKN DNTGYDFDSN1200
1201 YARVGRHGSF ILPVSKSVPT PSLLIEGSIV QRKNIKIE |
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
New Feature: Upload Your Own Microscopy Data
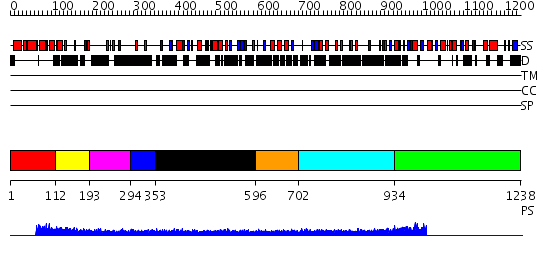
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..111] | N/A | No confident structure predictions are available. | |
| 2 | View Details | [112..192] | N/A | No confident structure predictions are available. | |
| 3 | View Details | [193..293] | N/A | Confident ab initio structure predictions are available. | |
| 4 | View Details | [294..352] | N/A | No confident structure predictions are available. | |
| 5 | View Details | [353..595] | N/A | No confident structure predictions are available. | |
| 6 | View Details | [596..701] | 1.08 | Solution structure of the Kruppel-associated box (KRAB) domain | |
| 7 | View Details | [702..933] | N/A | No confident structure predictions are available. | |
| 8 | View Details | [934..1238] | 1.46 | No description for 2i13A was found. |
Functions predicted (by domain):
| # | Gene Ontology predictions | |||||||||||||||
| 1 | No functions predicted. | |||||||||||||||
| 2 | No functions predicted. | |||||||||||||||
| 3 | No functions predicted. | |||||||||||||||
| 4 | No functions predicted. | |||||||||||||||
| 5 | No functions predicted. | |||||||||||||||
| 6 |
|
|||||||||||||||
| 7 | No functions predicted. | |||||||||||||||
| 8 | No functions predicted. |
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.89 |
Source: Reynolds et al. (2008)