













| General Information: |
|
| Name(s) found: |
mdj1 /
SPCC4G3.14
[Sanger Pombe]
|
| Description(s) found:
Found 20 descriptions. SHOW ALL |
|
| Organism: | Schizosaccharomyces pombe |
| Length: | 528 amino acids |
Gene Ontology: |
|
| Cellular Component: |
mitochondrion
[IDA mitochondrial inner membrane [ISS |
| Biological Process: |
proteolysis
[ISS protein folding [ISS |
| Molecular Function: |
zinc ion binding
[IEA]
unfolded protein binding [ISS Hsp70 protein binding [ISS |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MFSKYLQSRV CGLHSFTNSS AQQLFSKSIA HSSRRNFVIS SSCTKFRNVA IQRNAKREFS 60
61 RCAALKNFSY HARCFHATRA VWEMTDPYKT LGVSKSASAS EIKSAYYKLA KQYHPDANPD 120
121 KAAQDKFVEI KQAYEVLQDP KKKKAFDTYG AGAFKNGEFT GGDFEGFQNG FAGASSFSSG 180
181 FPGFNFEDLF GFSSRGPQAR RNTSFDVFVG EDIEASITID FMEAVRGAKK DLSYSVSSTC 240
241 SSCHGSGLQP GSHKSTCFAC KGTGQRLHFI PPSFHMQTTC DSCGGTGTTI PPNSACRSCM 300
301 GSGTVRERKT VSIDIPPGID DNTVLRVMGA GNDASTAKGG PNAKSRPGDL FATIHVRKHP 360
361 FFVREGTNVT YNAKIPMTTA ALGGTLRVPT LTGNVDLRVS PGTSTGDRIT MAGKGIRKVN 420
421 TSRYGNFYVN FEVTIPKILS PHERSLLEQL ADALNDSTAR RTQSSPSGTN SSTSTSSTSS 480
481 KHSTGISTEP TTGEENKQDG SVGGFFKRAF RRLHPDEDQN PKKDESSS |
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
New Feature: Upload Your Own Microscopy Data
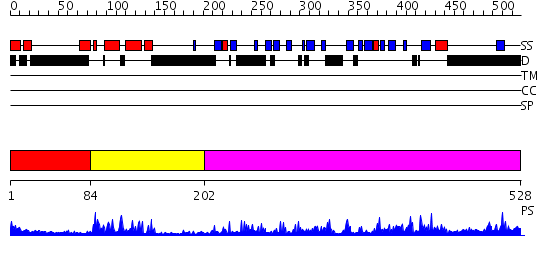
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..83] | N/A | No confident structure predictions are available. | |
| 2 | View Details | [84..201] | 23.0 | J-DOMAIN (RESIDUES 1-77) OF THE ESCHERICHIA COLI N-TERMINAL FRAGMENT (RESIDUES 1-104) OF THE MOLECULAR CHAPERONE DNAJ, NMR, 20 STRUCTURES | |
| 3 | View Details | [202..528] | 50.39794 | The crystal structure of Hsp40 Ydj1 |
Functions predicted (by domain):
| # | Gene Ontology predictions | |||||||||||||||||||||||||||||||||||||||||||||||||||
| 1 | No functions predicted. | |||||||||||||||||||||||||||||||||||||||||||||||||||
| 2 |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||
| 3 |
|
| Source: Reynolds et al. 2008. Manuscript submitted |

|