













| General Information: |
|
| Name(s) found: |
SPCC70.03c
[Sanger Pombe]
|
| Description(s) found:
Found 19 descriptions. SHOW ALL |
|
| Organism: | Schizosaccharomyces pombe |
| Length: | 492 amino acids |
Gene Ontology: |
|
| Cellular Component: |
mitochondrial matrix
[ISS |
| Biological Process: |
glutamate biosynthetic process
[ISS proline catabolic process [ISS |
| Molecular Function: |
proline dehydrogenase activity
[ISS |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MRAFRLASGV LRNRKVILGI GAGSLITAGN IKIRNDSKFD AFFAKGFPDE LQHRSLFSVL 60
61 RSAFVYEICS RAWLVKLSLG AMSLCDVFHL SFLYNPFCRY TFYKHFCGGE TPQAVMATMD 120
121 TLQAAGITSC LNYSREVDLD GDMDVNKIAS QGVVPPQVPV PSEKNQKVLR QIADKAFESN 180
181 MHIIDMATYK PGTVCAVKLT PFINPLVLQR YNSILNQYPV ESACNYLEHL KSPELSTYEV 240
241 SELKKFWEYA DKLCQFAKEK QIPLFIDAEQ TYFQDCMHAV TVDLMRKYNK EVAIVHNTYQ 300
301 LYLKKSRKIM DDHIKKCVAE GWLMGAKLVR GAYLNSEPRF LIHDTKAETD KDFDSAVEAI 360
361 IAAAAKFAPG DPASASDPIA SRKGKWGIMV ASHNKKTMFE SVNLAETKKV DFTKTSFYLA 420
421 QLLGMADDIT YALAYSQRNQ QPNFCIVKYV SCGPISEVLP YLVRRARENI DALDRCKEER 480
481 AYYRQALRRR IF |
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
New Feature: Upload Your Own Microscopy Data
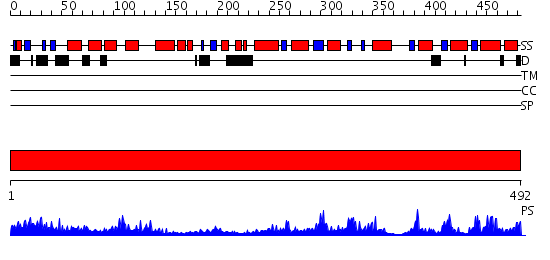
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..492] | 127.0 | Crystal structure of E. coli PutA proline dehydrogenase domain (residues 86-669) complexed with acetate |
Functions predicted (by domain):
| # | Gene Ontology predictions | |||||||||||||||||||||
| 1 |
|
| Source: Reynolds et al. 2008. Manuscript submitted |

|