













| General Information: |
|
| Name(s) found: |
gi|77553720
[NCBI NR]
|
| Description(s) found:
SHOW ONLY BEST |
|
| Organism: | Oryza sativa Japonica Group |
| Length: | 428 amino acids |
Gene Ontology: |
|
| Cellular Component: | NONE FOUND |
| Biological Process: | NONE FOUND |
| Molecular Function: | NONE FOUND |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MAAAGAGSSE VAARVLLQRY QPFAPPPGEY HQFGSGGAAA AGDMTEAVLI RTPLKRKHDR 60
61 EENEAAESND WMMSPGYTNP AGSPVPTPLS GKGSKAFAKS KAAKGQKSCP QTPLCASSPG 120
121 NPVTPVGGCR YDSSLGLLTK KFLNLLKGAP GGIVDLNNAA ETLEVQKRRI YDITNVLEGI 180
181 GLIEKKLKNN IRWKGIDDSR PGEVSDDMSI LQADIEALSL QEHSVDQQIS EMRDKLRGLT 240
241 EDENNQKWLY VTEDDIKSLP CFQNQTLIAI KAPHGTTLEV PDPDEVNDYP QRRYRIVLRS 300
301 TMGPIDVYLV SQFEEMSGME TPPRTVQPVS MDSLENPRTP LAAEPNKAAE SQPNIQDGLL 360
361 MPSDAPSSSQ DIGGMMKIVP SELDTDADYW LLSDAGVSIT DMWKTARILF QTVYHSCILP 420
421 FWLSSIIM |
SHOWING SINGLE HITS. [ Hide Single Hits ]
New Feature: Upload Your Own Microscopy Data
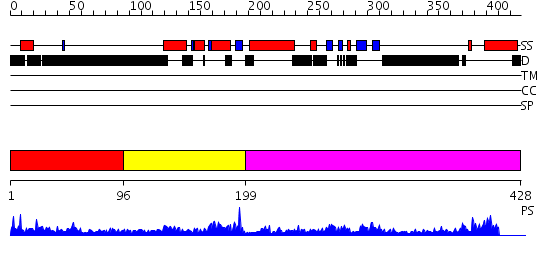
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..95] | 2.090994 | View MSA. No confident structure predictions are available. | |
| 2 | View Details | [96..198] | 22.09691 | Cell cycle transcription factor E2F-4 | |
| 3 | View Details | [199..428] | 28.69897 | Structure of the Rb C-terminal domain bound to an E2F1-DP1 heterodimer |
Functions predicted (by domain):
| # | Gene Ontology predictions | |||||||||||||||||||||||||||||||||
| 1 | No functions predicted. | |||||||||||||||||||||||||||||||||
| 2 |
|
|||||||||||||||||||||||||||||||||
| 3 | No functions predicted. |
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.88 |
Source: Reynolds et al. (2008)