













| General Information: |
|
| Name(s) found: |
ULP1D_ARATH
[Swiss-Prot]
|
| Description(s) found:
SHOW ONLY BEST |
|
| Organism: | Arabidopsis thaliana |
| Length: | 584 amino acids |
Gene Ontology: |
|
| Cellular Component: |
nucleus
[IDA]
|
| Biological Process: |
response to salt stress
[IGI]
protein desumoylation [IDA] proteolysis [ISS] vegetative to reproductive phase transition of meristem [IGI] |
| Molecular Function: |
SUMO-specific protease activity
[IDA]
cysteine-type peptidase activity [ISS] |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MTKRKKEVID VDCSEKKDFV IDWSSAMDKE DEVPELEIVN TTKPTPPPPP TFFSDDQTDS 60
61 PKLLTDRDLD EQLERKKAIL TLGPGLPDKG EKIRLKIADL EEEKQRRVLE GSKMEVDRSS 120
121 KVVSSTSSGS DVLPQGNAVS KDTSRGNADS KDTSRQGNAD SKEVSRSTFS AVFSKPKTDS 180
181 QSKKAFGKEL EDLGCERRKH KAGRKPVTRL SNGWRLLPDV GKAEHSAKQF DSGLKESKGN 240
241 KKSKEPYGKK RPMESSTYSL IDDDDDDDDD DDNDTSGHET PREWSWEKSP SQSSRRRKKS 300
301 EDTVINVDEE EAQPSTVAEQ AAELPEGLQE DICYPTRDDP HFVQVCLKDL ECLAPREYLT 360
361 SPVMNFYMRF LQQQISSSNQ ISADCHFFNT YFYKKLSDAV TYKGNDKDAF FVRFRRWWKG 420
421 IDLFRKAYIF IPIHEDLHWS LVIVCIPDKK DESGLTILHL DSLGLHSRKS IVENVKRFLK 480
481 DEWNYLNQDD YSLDLPISEK VWKNLPRRIS EAVVQVPQQK NDFDCGPFVL FFIKRFIEEA 540
541 PQRLKRKDLG MFDKKWFRPD EASALRIKIR NTLIELFRVS DQTE |
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
New Feature: Upload Your Own Microscopy Data
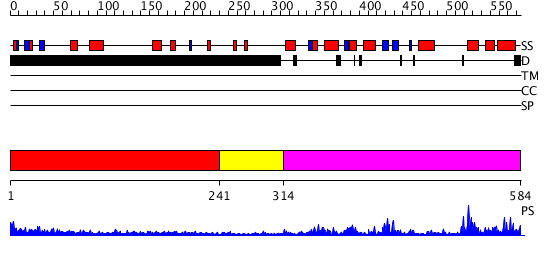
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..240] | 4.303996 | View MSA. No confident structure predictions are available. | |
| 2 | View Details | [241..313] | N/A | No confident structure predictions are available. | |
| 3 | View Details | [314..584] | 49.522879 | No description for 2iycA was found. |
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.99 |
Source: Reynolds et al. (2008)