













| General Information: |
|
| Name(s) found: |
gi|22328952
[NCBI NR]
gi|22136852 [NCBI NR] gi|17979390 [NCBI NR] |
| Description(s) found:
Found 9 descriptions. SHOW ALL |
|
| Organism: | Arabidopsis thaliana |
| Length: | 671 amino acids |
Gene Ontology: |
|
| Cellular Component: |
nucleolus
[IDA][ISS]
|
| Biological Process: |
rRNA processing
[IEA]
|
| Molecular Function: |
RNA binding
[IEA]
S-adenosylmethionine-dependent methyltransferase activity [IEA] |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MAALTRNKKK GSNSQTPPLN KQTKASPLKK AAKTQKPPLK KQRKCISEKK PLKKPEVSTD 60
61 EEEEEEENEQ SDEGSESGSD LFSDGDEEGN NDSDDDDDDD DDDDDDDEDA EPLAEDFLDG 120
121 SDNEEVTMGS DLDSDSGGSK LERKSRAIDR KRKKEVQDAD DEFKMNIKEK PDEFQLPTQK 180
181 ELEEEARRPP DLPSLQMRIR EIVRILSNFK DLKPKGDKHE RNDYVGQLKA DLSSYYGYNE 240
241 FLIGTLIEMF PVVELMELIE AFEKKRPTSI RTNTLKTRRR DLADILLNRG VNLDPLSKWS 300
301 KVGLIVYDSQ VPIGATPEYL AGFYMLQSAS SFLPVMALAP REKERVVDMA AAPGGKTTYV 360
361 AALMKNTGII YANEMKVPRL KSLSANLHRM GVTNTIVCNY DGRELTKVLG QSSVDRVLLD 420
421 APCSGTGVIS KDESVKTSKS ADDIKKFAHL QKQLILGAID LVDANSKTGG YIVYSTCSVM 480
481 IPENEAVIDY ALKNRDVKLV PCGLDFGRPG FSSFREHRFH PSLEKTRRFY PHVHNMDGFF 540
541 VAKLKKMSNA MQPSGNDEPA VTMEQAQVSS SDDDDEKAEA IEELEKPPVA SGQPKRESNT 600
601 KEDTNKRKNP RSKEIHKGKR NKNTKTESGN VEEPRKQKKK RSQWKNEIAQ AREEKRKTMR 660
661 ENAKETPKHR G |
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
New Feature: Upload Your Own Microscopy Data
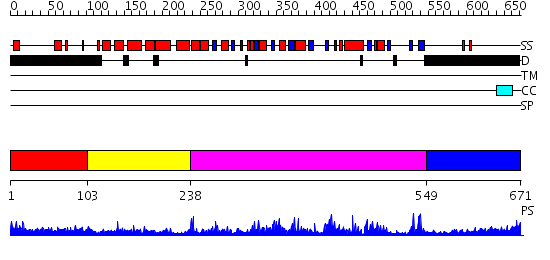
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..102] | 0.954 | Ribosomal protein L10 | |
| 2 | View Details | [103..237] | 5.30103 | Antitermination factor NusB | |
| 3 | View Details | [238..548] | 81.0 | Crystal Structure Analysis of Methyltransferase Homolog Protein from Pyrococcus Horikoshii | |
| 4 | View Details | [549..671] | 1.5 | No description for 2j63A was found. |
Functions predicted (by domain):
| # | Gene Ontology predictions | ||||||||||||||||||
| 1 | No functions predicted. | ||||||||||||||||||
| 2 |
|
||||||||||||||||||
| 3 |
|
||||||||||||||||||
| 4 | No functions predicted. |
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.99 |
Source: Reynolds et al. (2008)