













| General Information: |
|
| Name(s) found: |
gi|25054864
[NCBI NR]
gi|18417557 [NCBI NR] |
| Description(s) found:
Found 10 descriptions. SHOW ALL |
|
| Organism: | Arabidopsis thaliana |
| Length: | 1049 amino acids |
Gene Ontology: |
|
| Cellular Component: |
cellular_component
[ND]
|
| Biological Process: | NONE FOUND |
| Molecular Function: |
transition metal ion binding
[IEA]
oxidoreductase activity [IEA] acid phosphatase activity [IEA] |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MGVEEGAGVD KKITIGVCVM EKKVFSAPMG QIMDRIHAFG EFEIIHFGDK VILEDPVESW 60
61 PICDCLIAFY SSGYPLEKVQ AYSSLRKPFL VNELDPQYLL HDRRKVYEHL EMYGIPVPRY 120
121 ACVNRKVPDE DLDYFVEEED FVEVKGERFW KPFVEKPVNG DDHSIMIYYP SSAGGGMKEL 180
181 FRKVGNRSSE FHPDVRRVRR EGSYIYEEFM PTGGTDVKVY TVGPEYAHAE ARKSPVVDGV 240
241 VMRNPDGKEV RYPVLLTPAE KQMAREVCIA FRQAVCGFDL LRSEGSSYVC DVNGWSFVKN 300
301 SYKYYDDAAC VLRKMFLDAK APHLSSTIPP ILPWKINEPV QSNEGLTRQG SGIIGTFGQS 360
361 EELRCVIAIV RHGDRTPKQK VKLKVTEEKL LNLMLKYNGG KPRAETKLKT AVQLQDLLDA 420
421 TRMLIPRARS GESDSDAEDL EHADKLRQVK AVLEEGGHFS GIYRKVQLKP LKWVNVPKSD 480
481 GEGEEERPVE ALMVLKYGGV LTHAGRKQAE ELGRYFRNNM YPGEGTGLLR LHSTYRHDLK 540
541 IYSSDEGRVQ MSAAAFAKGL LDLEGQLTPI LVSLVSKDSS MLDGLDNASS EMEAAKAQLN 600
601 EIITAGSKMV HDHVSSELPW MTDGAGLPPH ADEHLPELVK LAKKVTEQVR LLAQDEHENL 660
661 AEPSAYDVVP PYDQAKALGK SNIDVGRIAA GLPCGSEGFL LMFARWRKLE RDLYNERRER 720
721 FDITQIPDVY DSCKYDLLHN SHLDLKGLDE LFKVAQLLAD GVIPNEYGIN PQQKLKIGSK 780
781 IARRLLGKIL IDLRNTREEA MSVAELKNSQ DQVSVSLYSS RKEDRYSQPK LFVKSDELRR 840
841 PSTGENKEED DDKETKYRLD PKYANVMTPE RHVRTRLYFT SESHIHSLMN VLRYCNLDES 900
901 LQGEESLVCQ SALDRLCKTK ELDYMSYVVL RLFENTEISL DDPKRFRIEL TFSRGADLSP 960
961 LEKKDEEAES LLREHTLPIM GPERLQEVGS CLTLETMEKM IRPFAMPAED FPPPCTPAGF1020
1021 SGYFSKSAAV LERLVKLWPF HKNTSNGKS |
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
New Feature: Upload Your Own Microscopy Data
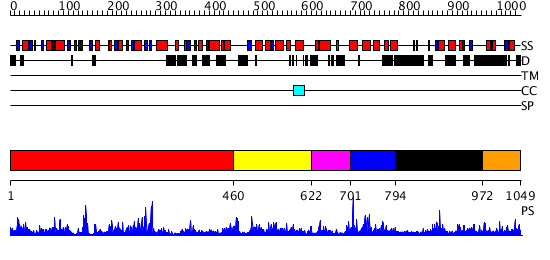
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..459] | 33.39794 | No description for 2dzdA was found. | |
| 2 | View Details | [460..621] | N/A | No confident structure predictions are available. | |
| 3 | View Details | [622..700] | N/A | No confident structure predictions are available. | |
| 4 | View Details | [701..793] | N/A | No confident structure predictions are available. | |
| 5 | View Details | [794..971] | 1.038971 | View MSA. No confident structure predictions are available. | |
| 6 | View Details | [972..1049] | N/A | No confident structure predictions are available. |
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.99 |
Source: Reynolds et al. (2008)