













| General Information: |
|
| Name(s) found: |
ANM16_ARATH
[Swiss-Prot]
|
| Description(s) found:
Found 9 descriptions. SHOW ALL |
|
| Organism: | Arabidopsis thaliana |
| Length: | 724 amino acids |
Gene Ontology: |
|
| Cellular Component: |
chloroplast
[IEA]
|
| Biological Process: |
protein amino acid methylation
[IEA]
|
| Molecular Function: |
methyltransferase activity
[IEA]
|
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MSPLSSLPPK TFISSFHCHS VTRLRRSVTA RTMSSQSSQR VFQLRQDPLT GNSEWIVIED 60
61 NDQPGTSTDG LLATTSYLDM LNDSRRNIAY RLAIEKTITE PCHVLDIGAG TGLLSMMAVR 120
121 AMRGDSKGMV TACESYLPMV KLMRKVMHKN GMTKNINLIN KRSDELKVGS EDIASRADVL 180
181 VSEILDSELL GEGLIPSLQH AHDMLLVDNP KTVPYRATTY CQLVESTFLC NLQDLRNNEA 240
241 KTSDGVRLVP PGLESLFGIK SQQYSMHVDA IEKEIKLLSE PVKIFEFDFW KRPESNGELD 300
301 VHIEAKTTGS VHAIISWWVL QLDSEGTIFY STAPRWIDSN SEIGVRDWCD HWKQCVWFTP 360
361 GTGVSISKGE KVHLHASHTC TNILYNLKKT QSLTHERTHF PLSTGDLHLT LPPERVAIYG 420
421 DSIYRQSLFE ATRKALQGKS YPQCLVIDDS LLLPLMALHI SNRSRVLSLS PGLQENAARY 480
481 FEAIADSNGF SKDRFEYFRD GKTNLAKAYP GKIDLLIGEP YYSGLENGLP WQNLRFWKDR 540
541 TLLDSVLSED AVVMPYKGVL RGCAMYLPDL WKSRCCLGSV EGFDHTLVNT TLGGCGDLPS 600
601 GKDSPCLPFF IWQCGETKIL SKEFTVMEFD FSKPITGPCS GEVQIEFIKP GVCHGIALWM 660
661 DWVMDEENST VISTGPDDKY WKQGVKLLGK PVTVRMEGPS SSIGIQASLD LSSNSELIVT 720
721 HTIS |
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
New Feature: Upload Your Own Microscopy Data
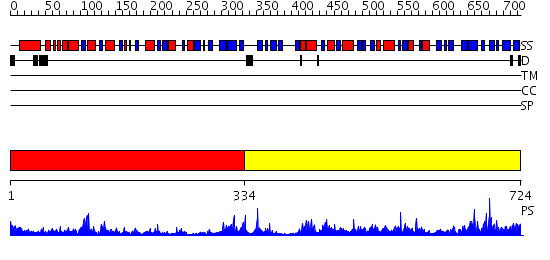
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..333] | 25.221849 | CmaA1 | |
| 2 | View Details | [334..724] | 90.69897 | Structure of the Predominant Protein Arginine Methyltransferase PRMT1 |
Functions predicted (by domain):
| # | Gene Ontology predictions | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 1 |
|
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 2 |
|
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.90 |
Source: Reynolds et al. (2008)