













| General Information: |
|
| Name(s) found: |
adl1 /
SPBC713.06
[Sanger Pombe]
|
| Description(s) found:
Found 17 descriptions. SHOW ALL |
|
| Organism: | Schizosaccharomyces pombe |
| Length: | 774 amino acids |
Gene Ontology: |
|
| Cellular Component: |
nucleus
[IDA |
| Biological Process: |
DNA repair
[IEA]
signal transduction [IEA] DNA replication [IEA] DNA recombination [IEA] |
| Molecular Function: |
DNA binding
[IEA]
ATP binding [IEA] DNA ligase (ATP) activity [IEA] signal transducer activity [IEA] |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MPPKKRMKNG SSLKSTSKKG EKSRNIITIQ DLFSKREAQL TDTPNKLLTD HDQSASDYAY 60
61 ALKLQQLFDS ENQATAPEKL PKDVIIPEEE YHTDTFNVVK ESNDKPKENL VTSEECKASF 120
121 FSTDSVNKDS TIDYDALQKD PLTYVKSCRA RFVSKDTKSF SYSSLANTFS LISSTKSRIR 180
181 IVTLLTNFLL TLLYADPDSL IATVWLCTNS IAPNFYGKNL GVGPAMYSKA LKEVCGITAS 240
241 ALKNLWNKYG DPGDVAFEAK VSVRTLSRPE PLTIKKVYST LLKIADSNGN GAQNRKLELT 300
301 KFLLISSNAE EVRYIGRSIM QNLRIGAVQN TMLASLSKAF FIFDNQNEIF NFNSDSLQQQ 360
361 FRQGEEIVKQ SFFQVPDYNI LVATLLREGI ENLKDNMSIR PGIPVKPMLG SITKNLQHML 420
421 ERLTDHNFSC EFKYDGQRAQ IHCDRLGNIK IFSRHLEEIT GRFPDVIEVA QLALKHSCDF 480
481 IIEGELVAID KSNGQILDFQ KLSTRERKKV TVADITIDVC VFVFDIMFCD GKSCLQMPLI 540
541 ERRRMFFEHF NLIPNRFQFV SSLETNEEQS IQEFFSLAIT NKCEGLMVKV LNGTNSKFPS 600
601 TYEPDKRGEG WIKVKQDYDD EFESLDLVPI GAWYGNGRKA GWFSPILLAV YNPDTGAYEA 660
661 VCKCMSGFSD QFYKELTQKY SLESGNSSLK PIYNFCETGK VTPQIYFAPQ EVWEIKGAQI 720
721 TSSPAYKAAL GLIQDDRGLS IRFPRFIRVR SDKGPEDAST NSILADMYMK QLNT |
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
New Feature: Upload Your Own Microscopy Data
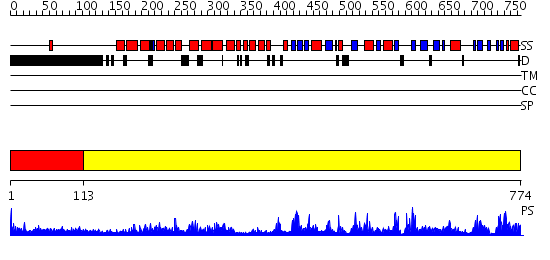
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..112] | N/A | No confident structure predictions are available. | |
| 2 | View Details | [113..774] | 1000.0 | Crystal Structure of Human DNA Ligase I bound to 5'-adenylated, nicked DNA |
Functions predicted (by domain):
| # | Gene Ontology predictions | |||||||||||||||||||||||||||||||||
| 1 | No functions predicted. | |||||||||||||||||||||||||||||||||
| 2 |
|
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.93 |
Source: Reynolds et al. (2008)