













| General Information: |
|
| Name(s) found: |
OSH7 /
YHR001W
[SGD]
|
| Description(s) found:
Found 24 descriptions. SHOW ALL |
|
| Organism: | Saccharomyces cerevisiae |
| Length: | 437 amino acids |
Gene Ontology: |
|
| Cellular Component: |
soluble fraction
[IDA cytoplasm [IDA |
| Biological Process: |
sterol transport
[IGI late endosome to vacuole transport [IMP endocytosis [IGI exocytosis [IGI sterol metabolic process [IGI |
| Molecular Function: |
oxysterol binding
[ISS |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MALNKLKNIP SLTNSSHSSI NGIASNAANS KPSGADTDDI DENDESGQSI LLNIISQLKP 60
61 GCDLSRITLP TFILEKKSML ERITNQLQFP DVLLEAHSNK DGLQRFVKVV AWYLAGWHIG 120
121 PRAVKKPLNP ILGEHFTAYW DLPNKQQAFY IAEQTSHHPP ESAYFYMIPE SNIRVDGVVV 180
181 PKSKFLGNSS AAMMEGLTVL QFLDIKDANG KPEKYTLSQP NVYARGILFG KMRIELGDHM 240
241 VIMGPKYQVD IEFKTKGFIS GTYDAIEGTI KDYDGKEYYQ ISGKWNDIMY IKDLREKSSK 300
301 KTVLFDTHQH FPLAPKVRPL EEQGEYESRR LWKKVTDALA VRDHEVATEE KFQIENRQRE 360
361 LAKKRAEDGV EFHSKLFRRA EPGEDLDYYI YKHIPEGTDK HEEQIRSILE TAPILPGQTF 420
421 TEKFSIPAYK KHGIQKN |
| PUBLICATION | TOPOLOGY | COCOMPLEXED PROTEINS | |
| View Details | Riffle et al. (2010) (Unpublished Data) |
|
OSH7 RPN10 RPN11 RPN12 RPN2 RPN3 RPN5 RPN6 RPN7 RPN8 RPN9 RPT2 RPT3 RPT4 RPT6 UBP6 |
| View Details | Ho Y, et al. (2002) |
|
CAF4 CCT2 CCT3 CCT5 CCT6 ENT2 OSH7 SRP54 TCP1 YBL029W |
| View Details | Qiu et al. (2008) |
|
OSH6 OSH7 |
The following runs contain data for this protein:
| BAIT | COMMENTS | PUBLICATION | |
| View Run | #32 Asynchronous Prep-No Phosphopeptide enrichment | Keck JM, et al. (2011) | |
| View Run | #29 Asynchronous Prep5-TiO2 Flowthrough | Keck JM, et al. (2011) |
SHOWING SINGLE HITS. [ Hide Single Hits ]
New Feature: Upload Your Own Microscopy Data
| PROTEIN(S) | PUBLICATION | |
| View Data |
|
Huh WK, et al. (2003) |
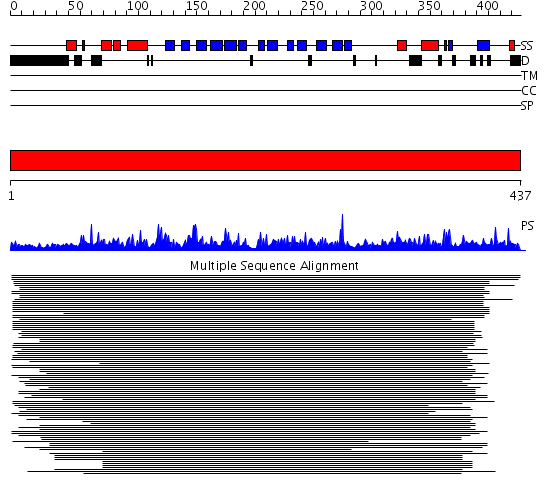
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..437] | 242.886057 | Oxysterol-binding protein No confident structure predictions are available. |
Functions predicted (by domain):
| # | Gene Ontology predictions | |||||||||||||||||||||||||||||||||
| 1 |
|
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.94 |
Source: Reynolds et al. (2008)