













| General Information: |
|
| Name(s) found: |
BNA6 /
YFR047C
[SGD]
|
| Description(s) found:
Found 25 descriptions. SHOW ALL |
|
| Organism: | Saccharomyces cerevisiae |
| Length: | 295 amino acids |
Gene Ontology: |
|
| Cellular Component: |
nucleus
[IDA cytoplasm [IDA |
| Biological Process: |
NAD biosynthetic process
[IDA |
| Molecular Function: |
nicotinate-nucleotide diphosphorylase (carboxylating) activity
[IDA |
| Sequence: |
|
| Sequence: [PDR BLAST] [ProtParam] |
1 11 21 31 41 51
| | | | | |
1 MPVYEHLLPV NGAWRQDVTN WLSEDVPSFD FGGYVVGSDL KEANLYCKQD GMLCGVPFAQ 60
61 EVFNQCELQV EWLFKEGSFL EPSKNDSGKI VVAKITGPAK NILLAERTAL NILSRSSGIA 120
121 TASHKIISLA RSTGYKGTIA GTRKTTPGLR RLEKYSMLVG GCDTHRYDLS SMVMLKDNHI 180
181 WATGSITNAV KNARAVCGFA VKIEVECLSE DEATEAIEAG ADVIMLDNFK GDGLKMCAQS 240
241 LKNKWNGKKH FLLECSGGLN LDNLEEYLCD DIDIYSTSSI HQGTPVIDFS LKLAH |
The following runs contain data for this protein:
| BAIT | COMMENTS | PUBLICATION | |
| View Run | #10 Mitotic Prep1-TiO2 Phosphopeptide enrichment | Keck JM, et al. (2011) |
NOT SHOWING SINGLE HITS. [ Show Single Hits ]
New Feature: Upload Your Own Microscopy Data
| PROTEIN(S) | PUBLICATION | |
| View Data |
|
Huh WK, et al. (2003) |
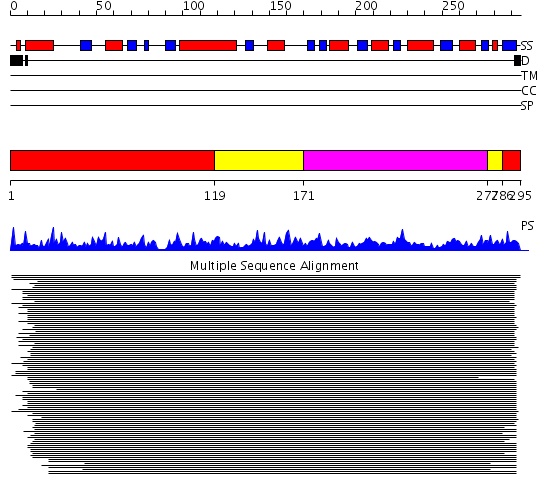
Domains predicted:
| # | Region(s) | Method | Confidence | Match Description | |
| 1 | View Details | [1..118] [286..295] |
398.69897 | Quinolinic acid phosphoribosyltransferase (Nicotinate-nucleotide pyrophosphorylase, NadC), C-terminal domain; Quinolinic acid phosphoribosyltransferase (Nicotinate-nucleotide pyrophosphorylase, NadC), N-terminal domain | |
| 2 | View Details | [119..170] [277..285] |
398.69897 | Quinolinic acid phosphoribosyltransferase (Nicotinate-nucleotide pyrophosphorylase, NadC), C-terminal domain; Quinolinic acid phosphoribosyltransferase (Nicotinate-nucleotide pyrophosphorylase, NadC), N-terminal domain | |
| 3 | View Details | [171..276] | 398.69897 | Quinolinic acid phosphoribosyltransferase (Nicotinate-nucleotide pyrophosphorylase, NadC), C-terminal domain; Quinolinic acid phosphoribosyltransferase (Nicotinate-nucleotide pyrophosphorylase, NadC), N-terminal domain |
Functions predicted (by domain):
| # | Gene Ontology predictions | ||||||||||||||||||
| 1 |
|
||||||||||||||||||
| 2 | No functions predicted. | ||||||||||||||||||
| 3 | No functions predicted. |
| Protein predicted to be: | GLOBULAR (No transmembrane regions or signal peptide) |
| Confidence of classification: | 0.99 |
Source: Reynolds et al. (2008)